Polypurine reverse-Hoogsteen hairpin
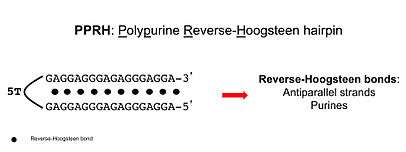
Polypurine reverse-Hoogsteen hairpins (PPRHs) are non-modified oligonucleotides formed by double-stranded DNA molecules containing two polypurine domains linked by a pentathymidine loop. The two domains are bound by intramolecular reverse-Hoogsteen bonds allowing the formation of a hairpin structure.
Properties

PPRHs can bind to polypyrimidine stretches in either single- or double stranded DNA by Watson and Crick bonds establishing triple-stranded DNA structures. The formation of PPRHs triplexes takes place at physiological pH. PPRHs provoke a strand displacement.[1] of the homopurine sequence of the target dsDNA, opening the two strands of the DNA. There are two types of PPRHs: i) Template-PPRHs[2] that bind to the template strand of DNA, inhibiting transcription; and ii) Coding-PPRHs[3] that bind to the coding strand of the DNA altering splicing. Both types of PPRHs decrease gene expression. PPRHs present high stability in serum and cells and show lack of immunogenicity not activating the innate inflammatory response.[4]
Applications
PPRHs are gene silencing tools acting by different mechanisms than triplex forming oligonucleotides (TFOs), antisense oligonucleotides or siRNAs. Upon binding to their targets, PPRHs can decrease the mRNA and protein levels of the selected genes. Their action has been demonstrated in vitro for a number of genes involved in metabolism (DHFR), proliferation (mTOR), DNA topology (TOP1), lifespan and senescence (telomerase), apoptosis (survivin, BCL2), transcription factors (MYC), and proto-oncogenes (MDM2)[5] as part of a cancer gene therapy strategy. Their preclinical proof of principle has been proven in vivo using the antiapoptotic survivin gene[6]
Design and improvements

PPRHs can be designed for virtually any gene in the genome by searching for polypirimidine stretches in the sequence of the desired gene. Optimal lengths for each domain of the PPRHs are within 20-30 nucleotides. The total length of a typical PPRH is 55 nucleotides considering two domains of 25 bases plus 5T for the linking loop. If purine interruptions are encountered (up to three) within the polypirimidine target, the highest affinity of PPRH binding is achieved by placing in the hairpin the complementary base (a pyrimidine) in front of the purines[7] (Wild type-PPRH).
Wedge-PPRH
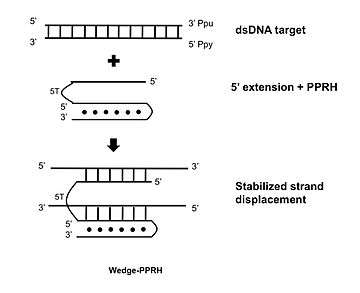
A further development consists in extending the 5' flank of the PPRH with a sequence complementary to the displaced polypurine strand of the target dsDNA which stabilizes the strand displacement, producing additional binding and functionality.[7]
References
- ↑ Coma, Silvia; Noe, Veronique; Eritja, Ramon; Ciudad, Carlos (2005). "Strand Displacement of Double-Stranded DNA by Triplex-Forming Antiparallel Purine-Hairpins". Oligonucleotides. 15: 269–83. PMID 16396621. doi:10.1089/oli.2005.15.269.
- ↑ de Almagro, Cristina; Coma, Silvia; Noe, Veronique; Ciudad, Carlos (2009). "Polypurine Hairpins Directed against the Template Strand of DNA Knock Down the Expression of Mammalian Genes". The Journal of Biological Chemistry. 284: 11579–89. PMC 2670163
 . PMID 19261618. doi:10.1074/jbc.M900981200.
. PMID 19261618. doi:10.1074/jbc.M900981200. - ↑ de Almagro, Cristina; Mencia, Nuria; Noe, Veronique; Ciudad, Carlos (2011). "Coding Polypurine Hairpins Cause Target-Induced Cell Death in Breast Cancer Cells". Human Gene Therapy. 22: 451–63. PMID 20942657. doi:10.1089/hum.2010.102.
- ↑ Villalobos, Xenia; Rodriguez, Laura; Prevot, Jeanne; Oleaga, Carlota; Ciudad, Carlos; Noe, Veronique (2013). "Stability and Immunogenicity Properties of the Gene-Silencing Polypurine Reverse Hoogsteen Hairpins". Molecular Pharmaceutics. 11: 254–64. PMID 24251728. doi:10.1021/mp400431f.
- ↑ Villalobos, Xenia; Rodriguez, Laura; Sole, Anna; Liberos, Carolina; Mencia, Nuria; Ciudad, Carlos; Noe, Veronique (2015). "Effect of Polypurine Reverse Hoogsteen Hairpins on Relevant Cancer Target Genes in Different Human Cell Lines". Nucleic Acid Therapeutics. 25: 198–208. PMID 26042602. doi:10.1089/nat.2015.0531.
- ↑ Rodriguez, Laura; Villalobos, Xenia; Dakhel, Sheila; Padilla, Laura; Hervas, Rosa; Hernandez, Jose Luis; Ciudad, Carlos; Noe, Veronique (2013). "Polypurine reverse Hoogsteen hairpins as a gene therapy tool against survivin in human prostate cancer PC3 cells in vitro and in vivo". Biochemical Pharmacology. 86: 1541–54. PMID 24070653. doi:10.1016/j.bcp.2013.09.013.
- 1 2 Rodriguez, Laura; Villalobos, Xenia; Sole, Anna; Liberos, Carolina; Ciudad, Carlos; Noe, Veronique (2015). "Improved Design of PPRHs for Gene Silencing". Molecular Pharmaceutics. 25: 198–208. PMID 26042602. doi:10.1089/nat.2015.0531.
External links
- Triplex-Forming Oligonucleotide Target Sequence Search Tool: A Searching Tool to find Polypurine and Polypyrimidine stretches in DNA