Sequence homology

Sequences are the amino acids for residues 120-180 of the proteins. Residues that are conserved across all sequences are highlighted in grey. Below the protein sequences is a key denoting conserved sequence (*), conservative mutations (:), semi-conservative mutations (.), and non-conservative mutations ( ).[1]
Sequence homology is the biological homology between protein or DNA sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry either because of a speciation event (orthologs), or because of a duplication event (paralogs).[2]
Homology among proteins or DNA is typically inferred from their sequence similarity. Significant similarity is strong evidence that two sequences are related by divergent evolution of a common ancestor. Alignments of multiple sequences are used to indicate which regions of each sequence are homologous.
The term "percent homology" is often used to mean "sequence similarity". The percentage of identical residues (percent identity) or the percentage of residues conserved with similar physicochemical properties (percent similarity), e.g. leucine and isoleucine, is usually used to "quantify the homology". Based on the definition of homology specified above this terminology is incorrect since sequence similarity is the observation, homology is the conclusion. Sequences are either homologous or not. As with anatomical structures, high sequence similarity might occur because of convergent evolution, or, as with shorter sequences, by chance, meaning that they are not homologous. Homologous sequence regions are also called conserved. This is not to be confused with conservation in amino acid sequences, where the amino acid at a specific position has been substituted with a different one that has functionally equivalent physicochemical properties.
Partial homology can occur where a segment of the compared sequences has a shared origin, while the rest does not. Such partial homology may result from a gene fusion event.
Orthology
Homologous sequences are orthologous if they are inferred to be descended from the same ancestral sequence separated by a speciation event: when a species diverges into two separate species, the copies of a single gene in the two resulting species are said to be orthologous. Orthologs, or orthologous genes, are genes in different species that originated by vertical descent from a single gene of the last common ancestor. The term "ortholog" was coined in 1970 by the molecular evolutionist Walter Fitch.[3]
For instance, the plant Flu regulatory protein is present both in Arabidopsis (multicellular higher plant) and Chlamydomonas (single cell green algae). The Chlamydomonas version is more complex: it crosses the membrane twice rather than once, contains additional domains and undergoes alternative splicing. However it can fully substitute the much simpler Arabidopsis protein, if transferred from algae to plant genome by means of genetic engineering. Significant sequence similarity and shared functional domains indicate that these two genes are orthologous genes,[4] inherited from the shared ancestor.
Orthology is strictly defined in terms of ancestry. Given that the exact ancestry of genes in different organisms is difficult to ascertain due to gene duplication and genome rearrangement events, the strongest evidence that two similar genes are orthologous is usually found by carrying out phylogenetic analysis of the gene lineage. Orthologs often, but not always, have the same function.[5]
Orthologous sequences provide useful information in taxonomic classification and phylogenetic studies of organisms. The pattern of genetic divergence can be used to trace the relatedness of organisms. Two organisms that are very closely related are likely to display very similar DNA sequences between two orthologs. Conversely, an organism that is further removed evolutionarily from another organism is likely to display a greater divergence in the sequence of the orthologs being studied.
Databases of orthologous genes
Given their tremendous importance for biology and bioinformatics, orthologous genes have been organized in several specialized databases that provide tools to identify and analyze orthologous gene sequences. These resources employ approaches that can be generally classified into those that use heuristic analysis of all pairwise sequence comparisons, and those that use phylogenetic methods. Sequence comparison methods were first pioneered in the COGs database in 1997.[6] These methods have been extended and automated in the following databases:
- eggNOG[7]
- InParanoid[8] focuses on pairwise ortholog relationships
- OrthoDB[9] appreciates that the orthology concept is relative to different speciation points by providing a hierarchy of orthologs along the species tree.
- OrthoMCL[10]
- OMA
- Roundup[11]
- OrthoMaM[12] for mammals
- OrthologID[13]
- GreenPhylDB[14] for plants
Tree-based phylogenetic approaches aim to distinguish speciation from gene duplication events by comparing gene trees with species trees, as implemented in databases such as
A third category of hybrid approaches uses both heuristic and phylogenetic methods to construct clusters and determine trees, for example
Paralogy
Homologous sequences are paralogous if they were created by a duplication event within the genome. For gene duplication events, if a gene in an organism is duplicated to occupy two different positions in the same genome, then the two copies are paralogous.
Paralogous genes often belong to the same species, but this is not necessary: for example, the hemoglobin gene of humans and the myoglobin gene of chimpanzees are paralogs. Paralogs can be split into in-paralogs (paralogous pairs that arose after a speciation event) and out-paralogs (paralogous pairs that arose before a speciation event). Between-species out-paralogs are pairs of paralogs that exist between two organisms due to duplication before speciation, whereas within-species out-paralogs are pairs of paralogs that exist in the same organism, but whose duplication event happened before speciation. Paralogs typically have the same or similar function, but sometimes do not: due to lack of the original selective pressure upon one copy of the duplicated gene, this copy is free to mutate and acquire new functions.
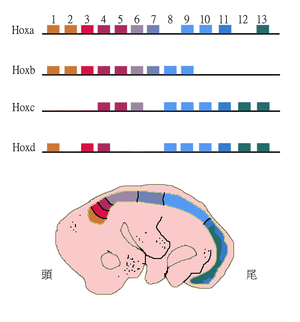
Paralogous genes can shape the structure of whole genomes and thus explain genome evolution to a large extent. Examples include the Homeobox (Hox) genes in animals. These genes not only underwent gene duplications within chromosomes but also whole genome duplications. As a result Hox genes in most vertebrates are clustered across multiple chromosomes with the HoxA-D clusters being the best studied.[20]
Another example are the globin genes which encode myoglobin and hemoglobin are considered to be ancient paralogs. Similarly, the four known classes of hemoglobins (hemoglobin A, hemoglobin A2, hemoglobin B, and hemoglobin F) are paralogs of each other. While each of these proteins serves the same basic function of oxygen transport, they have already diverged slightly in function: fetal hemoglobin (hemoglobin F) has a higher affinity for oxygen than adult hemoglobin. Function is not always conserved, however. Human angiogenin diverged from ribonuclease, for example, and while the two paralogs remain similar in tertiary structure, their functions within the cell are now quite different.
It is often asserted that orthologs are more functionally similar than paralogs of similar divergence, but several papers have challenged this notion.[21][22][23]
Regulation. Paralogs are often regulated differently, e.g. by having different tissue-specific expression patterns (see Hox genes). However, they can also be regulated differently on the protein level. For instance, Bacillus subtilis encodes two paralogues of glutamate dehydrogenase: GudB is constitutively transcribed whereas RocG is tightly regulated. In their active, oligomeric states, both enzymes show similar enzymatic rates. However, swaps of enzymes and promoters cause severe fitness losses, thus indicating promoter–enzyme coevolution. Characterization of the proteins shows that, compared to RocG, GudB's enzymatic activity is highly dependent on glutamate and pH.[24]
Paralogous regions
Sometimes, large chromosomal regions share gene content similar to other chromosomal regions within the same genome.[25] They are well characterised in the human genome, where they have been used as evidence to support the 2R hypothesis. Sets of duplicated, triplicated and quadruplicated genes, with the related genes on different chromosomes, are deduced to be remnants from genome or chromosomal duplications. A set of paralogy regions is together called a paralogon.[26] Well-studied sets of paralogy regions include regions of human chromosome 2, 7, 12 and 17 containing Hox gene clusters, collagen genes, keratin genes and other duplicated genes,[27] regions of human chromosomes 4, 5, 8 and 10 containing neuropeptide receptor genes, NK class homeobox genes and many more gene families,[28][29][30] and parts of human chromosomes 13, 4, 5 and X containing the ParaHox genes and their neighbors.[31] The Major histocompatibility complex (MHC) on human chromosome 6 has paralogy regions on chromosomes 1, 9 and 19.[32] Much of the human genome seems to be assignable to paralogy regions.[33]
Ohnology
Ohnologous genes are paralogous genes that have originated by a process of whole-genome duplication. The name was first given in honour of Susumu Ohno by Ken Wolfe.[34] Ohnologues are useful for evolutionary analysis because all ohnologues in a genome have been diverging for the same length of time (since their common origin in the whole genome duplication).
Xenology
Homologs resulting from horizontal gene transfer between two organisms are termed xenologs. Xenologs can have different functions, if the new environment is vastly different for the horizontally moving gene. In general, though, xenologs typically have similar function in both organisms. The term was coined by Walter Fitch.[3]
Gametology
Gametology denotes the relationship between homologous genes on non-recombining, opposite sex chromosomes. The term was coined by García-Moreno and Mindell.[35] 2000. Gametologs result from the origination of genetic sex determination and barriers to recombination between sex chromosomes. Examples of gametologs include CHDW and CHDZ in birds.[35]
See also
- Deep homology
- EggNOG (database)
- OrthoDB
- Orthologous MAtrix (OMA)
- Protein family
- Protein superfamily
- TreeFam
- Syntelog
References
- ↑ "Clustal FAQ #Symbols". Clustal. Retrieved 8 December 2014.
- ↑ Koonin EV (2005). "Orthologs, paralogs, and evolutionary genomics". Annual Review of Genetics. 39: 309–38. PMID 16285863. doi:10.1146/annurev.genet.39.073003.114725.
- 1 2 Fitch WM (June 1970). "Distinguishing homologous from analogous proteins". Systematic Zoology. 19 (2): 99–113. PMID 5449325. doi:10.2307/2412448.
- ↑ Falciatore A, Merendino L, Barneche F, et al. (January 2005). "The FLP proteins act as regulators of chlorophyll synthesis in response to light and plastid signals in Chlamydomonas". Genes & Development. 19 (1): 176–87. PMC 540235
 . PMID 15630026. doi:10.1101/gad.321305.
. PMID 15630026. doi:10.1101/gad.321305. - ↑ Fang G, Bhardwaj N, Robilotto R, Gerstein MB (March 2010). "Getting started in gene orthology and functional analysis". PLoS Computational Biology. 6 (3): e1000703. PMC 2845645
 . PMID 20361041. doi:10.1371/journal.pcbi.1000703.
. PMID 20361041. doi:10.1371/journal.pcbi.1000703. - ↑ COGs: Clusters of Orthologous Groups of proteins
Tatusov RL, Koonin EV, Lipman DJ (October 1997). "A genomic perspective on protein families". Science. 278 (5338): 631–7. PMID 9381173. doi:10.1126/science.278.5338.631. - ↑ eggNOG: evolutionary genealogy of genes: Non-supervised Orthologous Groups
Muller J, Szklarczyk D, Julien P, et al. (January 2010). "eggNOG v2.0: extending the evolutionary genealogy of genes with enhanced non-supervised orthologous groups, species and functional annotations". Nucleic Acids Research. 38 (Database issue): D190–5. PMC 2808932 . PMID 19900971. doi:10.1093/nar/gkp951.
. PMID 19900971. doi:10.1093/nar/gkp951. - ↑ Inparanoid: Eukaryotic Ortholog Groups
Ostlund G, Schmitt T, Forslund K, et al. (January 2010). "InParanoid 7: new algorithms and tools for eukaryotic orthology analysis". Nucleic Acids Research. 38 (Database issue): D196–203. PMC 2808972 . PMID 19892828. doi:10.1093/nar/gkp931.
. PMID 19892828. doi:10.1093/nar/gkp931. - ↑ Zdobnov, EM; Tegenfeldt, F; Kuznetsov, D; Waterhouse, RM; Simão, FA; Ioannidis, P; Seppey, M; Loetscher, A; Kriventseva, EV (28 November 2016). "OrthoDB v9.1: cataloging evolutionary and functional annotations for animal, fungal, plant, archaeal, bacterial and viral orthologs.". Nucleic Acids Research. 45: D744–D749. PMID 27899580. doi:10.1093/nar/gkw1119.
- ↑ OrthoMCL: Identification of Ortholog Groups for Eukaryotic Genomes
Chen F, Mackey AJ, Stoeckert CJ, Roos DS (January 2006). "OrthoMCL-DB: querying a comprehensive multi-species collection of ortholog groups". Nucleic Acids Research. 34 (Database issue): D363–8. PMC 1347485 . PMID 16381887. doi:10.1093/nar/gkj123.
. PMID 16381887. doi:10.1093/nar/gkj123. - ↑ Roundup
Deluca TF, Wu IH, Pu J, et al. (August 2006). "Roundup: a multi-genome repository of orthologs and evolutionary distances". Bioinformatics (Oxford, England). 22 (16): 2044–6. PMID 16777906. doi:10.1093/bioinformatics/btl286. - ↑ OrthoMaM
Ranwez V, Delsuc F, Ranwez S, Belkhir K, Tilak MK, Douzery EJ (2007). "OrthoMaM: a database of orthologous genomic markers for placental mammal phylogenetics". BMC Evolutionary Biology. 7: 241. PMC 2249597 . PMID 18053139. doi:10.1186/1471-2148-7-241.
. PMID 18053139. doi:10.1186/1471-2148-7-241. - ↑ OrthologID
Chiu JC, Lee EK, Egan MG, Sarkar IN, Coruzzi GM, DeSalle R (March 2006). "OrthologID: automation of genome-scale ortholog identification within a parsimony framework". Bioinformatics. 22 (6): 699–707. PMID 16410324. doi:10.1093/bioinformatics/btk040. - ↑ GreenPhylDB
Conte MG, Gaillard S, Lanau N, Rouard M, Périn C (January 2008). "GreenPhylDB: a database for plant comparative genomics". Nucleic Acids Research. 36 (Database issue): D991–8. PMC 2238940 . PMID 17986457. doi:10.1093/nar/gkm934.
. PMID 17986457. doi:10.1093/nar/gkm934. - ↑ TreeFam: Tree families database
Ruan J, Li H, Chen Z, et al. (January 2008). "TreeFam: 2008 Update". Nucleic Acids Research. 36 (Database issue): D735–40. PMC 2238856 . PMID 18056084. doi:10.1093/nar/gkm1005.
. PMID 18056084. doi:10.1093/nar/gkm1005. - ↑ TreeFam: Tree families database
van der Heijden RT, Snel B, van Noort V, Huynen MA (2007). "Orthology prediction at scalable resolution by phylogenetic tree analysis". BMC Bioinformatics. 8: 83. PMC 1838432 . PMID 17346331. doi:10.1186/1471-2105-8-83.
. PMID 17346331. doi:10.1186/1471-2105-8-83. - ↑ Fulton DL, Li YY, Laird MR, Horsman BG, Roche FM, Brinkman FS (2006). "Improving the specificity of high-throughput ortholog prediction". BMC Bioinformatics. 7: 270. PMC 1524997
 . PMID 16729895. doi:10.1186/1471-2105-7-270.
. PMID 16729895. doi:10.1186/1471-2105-7-270. - ↑ Vilella AJ, Severin J, Ureta-Vidal A, Heng L, Durbin R, Birney E (February 2009). "EnsemblCompara GeneTrees: Complete, duplication-aware phylogenetic trees in vertebrates". Genome Research. 19 (2): 327–35. PMC 2652215
 . PMID 19029536. doi:10.1101/gr.073585.107.
. PMID 19029536. doi:10.1101/gr.073585.107. - ↑ Sayers EW, Barrett T, Benson DA, et al. (January 2011). "Database resources of the National Center for Biotechnology Information". Nucleic Acids Research. 39 (Database issue): D38–51. PMC 3013733
 . PMID 21097890. doi:10.1093/nar/gkq1172.
. PMID 21097890. doi:10.1093/nar/gkq1172. - 1 2 Zakany, Jozsef; Duboule, Denis (2007-08-01). "The role of Hox genes during vertebrate limb development". Current Opinion in Genetics & Development. 17 (4): 359–366. ISSN 0959-437X. PMID 17644373. doi:10.1016/j.gde.2007.05.011.
- ↑ Studer, R.A.; Robinson-Rechavi, M. (May 2009). "How confident can we be that orthologs are similar, but paralogs differ?". Trends in Genetics. 25 (5): 210–6. PMID 19368988. doi:10.1016/j.tig.2009.03.004.
- ↑ Nehrt NL, Clark WT, Radivojac P, Hahn MW (June 2011). "Testing the ortholog conjecture with comparative functional genomic data from mammals". PLoS Computational Biology. 7 (6): e1002073. PMC 3111532
 . PMID 21695233. doi:10.1371/journal.pcbi.1002073.
. PMID 21695233. doi:10.1371/journal.pcbi.1002073. - ↑ Eisen, Jonathan. "Special Guest Post & Discussion Invitation from Matthew Hahn on Ortholog Conjecture Paper".
- ↑ Noda‐Garcia, Lianet; Romero, Maria Luisa Romero; Longo, Liam M.; Kolodkin‐Gal, Ilana; Tawfik, Dan S. (2017-07-01). "Bacilli glutamate dehydrogenases diverged via coevolution of transcription and enzyme regulation". EMBO reports. 18 (7): 1139–1149. ISSN 1469-221X. PMID 28468957. doi:10.15252/embr.201743990.
- ↑ Lundin, L.G. (1993). "Evolution of the vertebrate genome as reflected in paralogous chromosomal regions in man and the house mouse". Genomics. 16 (1): 1–19. PMID 8486346. doi:10.1006/geno.1993.1133.
- ↑ Coulier, F; Popovici, C; Villet, R; Birnbaum, D (Dec 15, 2000). "MetaHox gene clusters". The Journal of experimental zoology. 288 (4): 345–51. PMID 11144283. doi:10.1002/1097-010X(20001215)288:4<345::AID-JEZ7>3.0.CO;2-Y.
- ↑ Ruddle, FH; Bentley, KL; Murtha, M.T.; Risch, N. (1994). "Gene loss and gain in the evolution of the vertebrates". Development (Cambridge, England). Supplement: 155–61. PMID 7579516.
- ↑ Pébusque, M.J.; Coulier, F.; Birnbaum, D.; Pontarotti, P. (September 1998). "Ancient large-scale genome duplications: phylogenetic and linkage analyses shed light on chordate genome evolution". Molecular Biology and Evolution. 15 (9): 1145–59. PMID 9729879. doi:10.1093/oxfordjournals.molbev.a026022.
- ↑ Larsson, T.A.; Olsson, F.; Sundstrom, .; Lundin, L.G.; Brenner, S.; Venkatesh, B.; Larhammar, D. (Jun 25, 2008). "Early vertebrate chromosome duplications and the evolution of the neuropeptide Y receptor gene regions.". BMC Evolutionary Biology. 8: 184. PMC 2453138
 . PMID 18578868. doi:10.1186/1471-2148-8-184.
. PMID 18578868. doi:10.1186/1471-2148-8-184. - ↑ Pollard, Sophie L.; Holland, Peter W.H. (2000). "Evidence for 14 homeobox gene clusters in human genome ancestry". Current Biology. 10 (17): 1059–1062. PMID 10996074. doi:10.1016/S0960-9822(00)00676-X.
- ↑ Mulley, J.F.; Chiu, C.H.; Holland, P.W. (2006). "Breakup of a homeobox cluster after genome duplication in teleosts". Proceedings of the National Academy of Sciences of the United States of America. 103 (27): 10369–72. PMC 1502464
 . PMID 16801555. doi:10.1073/pnas.0600341103.
. PMID 16801555. doi:10.1073/pnas.0600341103. - ↑ Flajnik, Martin F.; Kasahara, Masanori (2001). "Comparative Genomics of the MHC". Immunity. 15 (3): 351–362. PMID 11567626. doi:10.1016/S1074-7613(01)00198-4.
- ↑ McLysaght, Aoife; Hokamp, Karsten; Wolfe, Kenneth H. (2002). "Extensive genomic duplication during early chordate evolution". Nature Genetics. 31 (2): 200–204. PMID 12032567. doi:10.1038/ng884.
- ↑ Wolfe K (May 2000). "Robustness--it's not where you think it is". Nature Genetics. 25 (1): 3–4. PMID 10802639. doi:10.1038/75560.
- 1 2 García-Moreno, Jaime; Mindell, David P. (2000-12-01). "Rooting a Phylogeny with Homologous Genes on Opposite Sex Chromosomes (Gametologs): A Case Study Using Avian CHD". Molecular Biology and Evolution. 17 (12): 1826–1832. ISSN 0737-4038. doi:10.1093/oxfordjournals.molbev.a026283.