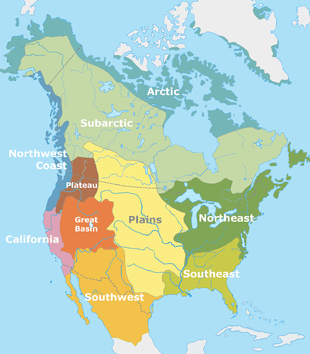
Genetic history of indigenous peoples of the Americas

The genetic history of indigenous peoples of the Americas primarily focuses on Human Y-chromosome DNA haplogroups and Human mitochondrial DNA haplogroups.[1] Autosomal "atDNA" markers are also used, but differ from mtDNA or Y-DNA in that they overlap significantly.[2] The genetic pattern indicates Indigenous Amerindians experienced two very distinctive genetic episodes; first with the initial peopling of the Americas, and secondly with European colonization of the Americas.[3][4] The former is the determinant factor for the number of gene lineages, zygosity mutations and founding haplotypes present in today's Indigenous Amerindian populations.[5]
Analyses of genetics among Amerindian and Siberian populations have been used to argue for early isolation of founding populations on Beringia[6] and for later, more rapid migration from Siberia through Beringia into the New World.[7] The microsatellite diversity and distributions of the Y lineage specific to South America indicates that certain Amerindian populations have been isolated since the initial colonization of the region.[8] The Na-Dené, Inuit and Indigenous Alaskan populations exhibit Haplogroup Q-M242; however, they are distinct from other indigenous Amerindians with various mtDNA and atDNA mutations.[9][10][11] This suggests that the peoples who first settled the northern extremes of North America and Greenland derived from later migrant populations than those who penetrated farther south in the Americas.[12][13] Linguists and biologists have reached a similar conclusion based on analysis of Amerindian language groups and ABO blood group system distributions.[14][15][16]
There is general agreement among anthropologists that the source populations for the migration into the Americas originated from an area somewhere east of the Yenisei River.[17] The common occurrence of the mtDNA Haplogroups A, B, C, and D among eastern Asian and Amerindian populations has long been recognized, along with the presence of Haplogroup X.[18] As a whole, the greatest frequency of the four Amerindian associated haplogroups occurs in the Altai-Baikal region of southern Siberia.[19] Some subclades of C and D closer to the Amerindian subclades occur among Mongolian, Amur, Japanese, Korean, and Ainu populations.[18][20]
Background

The Y chromosome is passed down directly from father to son; all male humans (Y chromosomes) today trace back to a single prehistoric father termed "Y-chromosomal Adam" originating in Africa.[21] The Y chromosome spans about 60 million base pairs (the building blocks of DNA) and represents about 2 percent of the total DNA in all human cells.[22] The original "Y chromosomal Adam"-DNA sequencing has mutated rarely over the 20,000 generations, but each time a new mutation occurs, there is a new branch in a haplogroup, resulting in a new subclade (single-nucleotide polymorphism (SNP)).[23]
Both females and males inherit their Mitochondrial DNA (mtDNA) from only their mother.[24] MtDNA mutations are also passed down relatively unchanged from generation to generation, so all humans share the same mtDNA-types. The logical extension of this is that all humans ultimately trace back to one woman, who is commonly referred to as Mitochondrial Eve.[25][26] This line of biological inheritance, therefore, stops with each male.[27] Consequently, Y-DNA is more commonly used by the general public for tracing genetic heritage of a direct male line.[27][28][29]
An autosome (atDNA) is a chromosome that is not a sex chromosome – that is to say, there are an equal number of copies of the chromosome in males and females.[2] Autosomal DNA testing is generally used to determine the "genetic percentages" of a person's ancestry from particular continents/regions or to identify the countries and "tribes" of origin on an overall basis. Genetic admixture tests arrive at these percentages by examining locations (SNPs) on the DNA where one nucleotide has "mutated" or "switched" to a different nucleotide.[2] One way to examine the support for particular colonization routes within the American landmass is to determine whether a closer relationship between zygosity and geography is observed when "effective" geographic distances are computed along these routes, rather than along shortest-distance paths.[30]
Y-DNA
The Y chromosome consortium has established a system of defining Y-DNA haplogroups by letters A through to T, with further subdivisions using numbers and lower-case letters.[31]
Haplogroup Q
Q-M242 (mutational name) is the defining (SNP) of Haplogroup Q (Y-DNA) (phylogenetic name). Within the Q clade, there are 14 haplogroups marked by 17 SNPs.2009[32][33] In Eurasia, haplogroup Q is found among indigenous Siberian populations, such as the modern Chukchi and Koryak peoples. In particular, two groups exhibit large concentrations of the Q-M242 mutation, the Ket (93.8%) and the Selkup (66.4%) peoples.[34] The Ket are thought to be the only survivors of ancient wanderers living in Siberia.[35] Their population size is very small; there are fewer than 1,500 Ket in Russia.2002[35] The Selkup have a slightly larger population size than the Ket, with approximately 4,250 individuals.[36]
Starting the Paleo-Indians period, a migration to the Americas across the Bering Strait (Beringia) by a small population carrying the Q-M242 mutation took place.[37] A member of this initial population underwent a mutation, which defines its descendant population, known by the Q-M3 (SNP) mutation.[38] These descendants migrated all over the Americas.[32]
Haplogroup Q-M3 is defined by the presence of the rs3894 (M3) (SNP).[3][35][39] The Q-M3 mutation is roughly 15,000 years old as that is when the initial migration of Paleo-Indians into the Americas occurred.[40][41] Q-M3 is the predominant haplotype in the Americas, at a rate of 83% in South American populations,[42] 50% in the Na-Dené populations, and in North American Eskimo-Aleut populations at about 46%.[34] With minimal back-migration of Q-M3 in Eurasia, the mutation likely evolved in east-Beringia, or more specifically the Seward Peninsula or western Alaskan interior. The Beringia land mass began submerging, cutting off land routes.[34][43][44]
Since the discovery of Q-M3, several subclades of M3-bearing populations have been discovered. An example is in South America, where some populations have a high prevalence of (SNP) M19 which defines subclade Q-M19.[42] M19 has been detected in (59%) of Amazonian Ticuna men and in (10%) of Wayuu men.[42] Subclade M19 appears to be unique to South American Indigenous peoples, arising 5,000 to 10,000 years ago.[42]

This suggests that population isolation and perhaps even the establishment of tribal groups began soon after migration into the South American areas.[35][46] Other American subclades include Q-L54, Q-Z780, Q-MEH2, Q-SA01, and Q-M346 lineages. In Canada, two other lineages have been found. These are Q-P89.1 and Q-NWT01.
The principal-component analysis suggests a close genetic relatedness between some North American Amerindians (the Chipewyan and the Cheyenne) and certain populations of central/southern Siberia (particularly the Kets, Yakuts, Selkups, and Altays), at the resolution of major Y-chromosome haplogroups.[47] This pattern agrees with the distribution of mtDNA haplogroup X, which is found in North America, is absent from eastern Siberia, but is present in the Altais of southern central Siberia. Similarly, the Asian populations closest to Native Americans are characterized by a predominance of lineage P-M45* and low frequencies of C-RPS4Y.[47]
Haplogroup R1
Haplogroup R1 (Y-DNA) (specially R1b) is the second most predominant Y haplotype found among indigenous Amerindians after Q (Y-DNA).[48] The distribution of R1 is believed by some to be associated with the re-settlement of Eurasia following the last glacial maximum. One theory put forth is that R1 entered the Americas with the initial founding population,[49] suggesting prehistoric Amerindian immigration from Asia through Beringia[50][51] and correlating mostly with the frequency of haplogroups Q-M3 and P-M45*.[47] A second theory is that it was introduced during European colonization.[48] R1 is very common throughout all of Eurasia except East Asia and Southeast Asia. R1 (M173) is found predominantly in North American groups like the Ojibwe (50-79%), Seminole (50%), Sioux (50%), Cherokee (47%), Dogrib (40%) and Tohono O'odham (Papago) (38%).[48]
A study of Raghavan et al. 2013 found that autosomal evidence indicates that skeletal remain of a south-central Mongoloid Siberian child carrying R* y-dna (Mal'ta boy-1) "is basal to modern-day western Eurasians and genetically closely related to modern-day Amerindians, with no close affinity to east Asians." Sequencing of another south-central Siberian (Afontova Gora-2) revealed that "Siberian genetic signatures in modern-day Amerindians derive from a mixed ancestry of the First Americans."[51] It is further theorized if "Mal'ta might be a missing link, a representative of the Asian population that admixed both into proto Indo-Europeans and Native Americans."[52]
Haplogroup C-P39

Haplogroup C-M217 is mainly found in indigenous Siberians, Mongolians and Kazakhs. Haplogroup C-M217 is the most widespread and frequently occurring branch of the greater (Y-DNA) haplogroup C-M130. Haplogroup C-M217 descendant C-P39 is commonly found in today's Na-Dené speakers, with the highest frequency found among the Athabaskans at 42%.[37] This distinct and isolated branch C-P39 includes almost all the Haplogroup C-M217 Y-chromosomes found among all indigenous peoples of the Americas.[53] The Na-Dené groups are also unusual among indigenous peoples of the Americas in having a relatively high frequency of Q-M242 (25%).[34]
Some researchers feel that this may indicate that the Na-Dené migration occurred from the Russian Far East after the initial Paleo-Indian colonization, but prior to modern Inuit, Inupiat and Yupik expansions.[9][10][54]
mtDNA
Mitochondrial Eve is defined as the woman who was the matrilineal most recent common ancestor for all living humans. Mitochondrial Eve is estimated to have lived between 140,000 and 200,000 years ago,[55][56] Mitochondrial Eve is the most recent common matrilineal ancestor, not the most recent common ancestor.[57][58]

When studying human mitochondrial DNA (mtDNA) haplogroups, the results indicated until recently that Indigenous Amerindian haplogroups, including haplogroup X, are part of a single founding east Asian population. It also indicates that the distribution of mtDNA haplogroups and the levels of sequence divergence among linguistically similar groups were the result of multiple preceding migrations from Bering Straits populations.[24][59] All Indigenous Amerindian mtDNA can be traced back to five haplogroups, A, B, C, D and X.[60] More specifically, indigenous Amerindian mtDNA belongs to sub-haplogroups A2, B2, C1, D1, and X2a (with minor groups C4c, D2, D3, and D4h3).[6][59] This suggests that 95% of Indigenous Amerindian mtDNA is descended from a minimal genetic founding female population, comprising sub-haplogroups A2, B2, C1b, C1c, C1d, and D1.[60] The remaining 5% is composed of the X2a, D2, D3, C4, and D4h3 sub-haplogroups.[59][60]
X is one of the five mtDNA haplogroups found in Indigenous Amerindian peoples. Unlike the four main American mtDNA haplogroups (A, B, C and D), X is not at all strongly associated with east Asia.[35] Haplogroup X genetic sequences diverged about 20,000 to 30,000 years ago to give two sub-groups, X1 and X2. X2's subclade X2a occurs only at a frequency of about 3% for the total current indigenous population of the Americas.[35] However, X2a is a major mtDNA subclade in North America; among the Algonquian peoples, it comprises up to 25% of mtDNA types.[61][62] It is also present in lower percentages to the west and south of this area — among the Sioux (15%), the Nuu-chah-nulth (11%–13%), the Navajo (7%), and the Yakama (5%).[63] Haplogroup X is more strongly present in the Near East, the Caucasus, and Mediterranean Europe.[63] The predominant theory for sub-haplogroup X2a's appearance in North America is migration along with A, B, C, and D mtDNA groups, from a source in the Altai Mountains of central Asia.[64][65][66][67]
Sequencing of the mitochondrial genome from Paleo-Eskimo remains (3,500 years old) are distinct from modern Amerindians, falling within sub-haplogroup D2a1, a group observed among today's Aleutian Islanders, the Aleut and Siberian Yupik populations.[68] This suggests that the colonizers of the far north, and subsequently Greenland, originated from later coastal populations.[68] Then a genetic exchange in the northern extremes introduced by the Thule people (proto-Inuit) approximately 800–1,000 years ago began.[11][69] These final Pre-Columbian migrants introduced haplogroups A2a and A2b to the existing Paleo-Eskimo populations of Canada and Greenland, culminating in the modern Inuit.[11][69]

A 2013 study in Nature reported that DNA found in the 24,000-year-old remains of a young boy from the archaeological Mal'ta-Buret' culture suggest that up to one-third of the indigenous Americans may have ancestry that can be traced back to western Eurasians, who may have "had a more north-easterly distribution 24,000 years ago than commonly thought"[51] "We estimate that 14 to 38 percent of Amerindian ancestry may originate through gene flow from this ancient population," the authors wrote. Professor Kelly Graf said,
"Our findings are significant at two levels. First, it shows that Upper Paleolithic Siberians came from a cosmopolitan population of early modern humans that spread out of Africa to Europe and Central and South Asia. Second, Paleoindian skeletons like Buhl Woman with phenotypic traits atypical of modern-day indigenous Americans can be explained as having a direct historical connection to Upper Paleolithic Siberia."[70]
A route through Beringia is seen as more likely than the Solutrean hypothesis.[71] An abstract in a 2012 issue of the "American Journal of Physical Anthropology" states that "The similarities in ages and geographical distributions for C4c and the previously analyzed X2a lineage provide support to the scenario of a dual origin for Paleo-Indians. Taking into account that C4c is deeply rooted in the Asian portion of the mtDNA phylogeny and is indubitably of Asian origin, the finding that C4c and X2a are characterized by parallel genetic histories definitively dismisses the controversial hypothesis of an Atlantic glacial entry route into North America."[72]
Another study, also focused on the mtDNA (that which is inherited through only the maternal line),[73] revealed that the indigenous people of the Americas have their maternal ancestry traced back to a few founding lineages from East Asia, which would have arrived via the Bering strait. According to this study, it is probable that the ancestors of the Native Americans would have remained for a time in the region of the Bering Strait, after which there would have been a rapid movement of settling of the Americas, taking the founding lineages to South America.
According to a 2016 study, focused on mtDNA lineages, "a small population entered the Americas via a coastal route around 16.0 ka, following previous isolation in eastern Beringia for ~2.4 to 9 thousand years after separation from eastern Siberian populations. Following a rapid movement throughout the Americas, limited gene flow in South America resulted in a marked phylogeographic structure of populations, which persisted through time. All of the ancient mitochondrial lineages detected in this study were absent from modern data sets, suggesting a high extinction rate. To investigate this further, we applied a novel principal components multiple logistic regression test to Bayesian serial coalescent simulations. The analysis supported a scenario in which European colonization caused a substantial loss of pre-Columbian lineages".[74]
By autosomal DNA
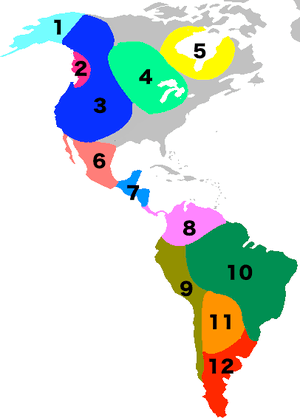
Genetic diversity and population structure in the American landmass is also done using autosomal (atDNA) micro-satellite markers genotyped; sampled from North, Central, and South America and analyzed against similar data available from other indigenous populations worldwide.[30][75] The Amerindian populations show a lower genetic diversity than populations from other continental regions.[75] Observed is a decreasing genetic diversity as geographic distance from the Bering Strait occurs, as well as a decreasing genetic similarity to Siberian populations from Alaska (the genetic entry point).[30][75]
Also observed is evidence of a higher level of diversity and lower level of population structure in western South America compared to eastern South America.[30][75] There is a relative lack of differentiation between Mesoamerican and Andean populations, a scenario that implies that coastal routes were easier for migrating peoples (more genetic contributors) to traverse in comparison with inland routes.[30]
The over-all pattern that is emerging suggests that the Americas were colonized by a small number of individuals (effective size of about 70), which grew by a factor of 10 over 800 – 1000 years.[35][76] The data also shows that there have been genetic exchanges between Asia, the Arctic, and Greenland since the initial peopling of the Americas.[76][77]
In 2014, the autosomal DNA of a 12,500+-year-old infant from Montana was sequenced.[78] The DNA was taken from a skeleton referred to as Anzick-1, found in close association with several Clovis artifacts. Comparisons showed strong affinities with DNA from Siberian sites, and virtually ruled out that particular individual had any close affinity with European sources (the "Solutrean hypothesis"). The DNA also showed strong affinities with all existing Amerindian populations, which indicated that all of them derive from an ancient population that lived in or near Siberia, the Upper Palaeolithic Mal'ta population.[79]
According to an autosomal genetic study from 2012,[80] Native Americans descend of at least three main migrant waves from East Asia. Most of it is traced back to a single ancestral population, called 'First Americans'. However, those who speak Inuit languages from the Arctic inherited almost half of their ancestry from a second East Asian migrant wave. And those who speak Na-dene, on the other hand, inherited a tenth of their ancestry from a third migrant wave. The initial settling of the Americas was followed by a rapid expansion southwards, by the coast, with little gene flow later, especially in South America. One exception to this are the Chibcha speakers, whose ancestry comes from both North and South America. [80]
Linguistic studies have backed up genetic studies, with ancient patterns having been found between the languages spoken in Siberia and those spoken in the Americas.[81]
Two 2015 autosomal DNA genetic studies confirmed the Siberian origins of the Natives of the Americas. However an ancient signal of shared ancestry with the Natives of Australia and Melanesia was detected among the Natives of the Amazon region. The migration coming out of Siberia would have happened 23000 years ago.[82][83]
Overlaps between DNA types
Populations that have a specific combination of autosome, Y and MT-haplogroup mutations can generally be found with regional variations. Autosomes, Y mutations and mt mutations do not necessarily occur at a similar time and there are differential rates of sexual selection between the two sex chromosomes.[84] This combined with population bottlenecks, the founder effect, mitochondrial mutations and genetic drift will alter the genetic composition of isolated populations, resulting in very distinguishable mutation patterns.[85] (i.e. Taínos,[86] Fuegians,[87] Inuit,[88] Yupik[89] and Algonquian[35])
The rough overlaps between Y-DNA and mtDNA between the Americas, Circumpolar north, and Siberian indigenous populations are:
| Y-DNA haplogroup(s) - | mtDNA haplogroup(s) - | Geographical area(s) |
|---|---|---|
| Q-M242, R1, C-M217 | A, X, C, D (N types), (M types) |
Russian far east, Americas, Arctic |
Old world genetic admixture

Interracial marriage and interracial sex and, more generally, the process of racial admixture, has its origins in prehistory. Racial mixing became widespread during European colonialism in the Age of Discovery.[90] Genetic exchange between two populations reduces the genetic distance between the populations and is measurable in DNA patterns.[84] During the Age of Discovery, beginning in the late 15th century, European explorers sailed the oceans, eventually reaching all the major continents.[91] During this time Europeans contacted many populations, some of which had been relatively isolated for millennia.[92] The genetic demographic composition of the Eastern Hemisphere has not changed significantly since the Age of Discovery. However, genetic demographics in the Western Hemisphere were radically altered by events following the voyages of Christopher Columbus.[92] The European colonization of the Americas brought contact between peoples of Europe, Africa and Asia and the Amerindian populations. As a result, the Americas today have significant and complex multiracial populations.[92] Many individuals who self-identify as one race exhibit genetic evidence of a multiracial ancestry.[93]
The European conquest of Latin America beginning in the late 15th century, was initially executed by male soldiers and sailors from the Iberian Peninsula (Spain and Portugal).[94] The new soldier-settlers fathered children with Amerindian women and later with African slaves.[95] These mixed-race children were generally identified by the Spanish colonist and Portuguese colonist as "Castas".[96] The subsequent North American fur trade during the 16th century brought many more European men, from France, Ireland, and Great Britain, who took North Amerindian women as wives.[97] Their children became known as "Métis" or "Bois-Brûlés" by the French colonists and "mixed-bloods", "half-breeds" or "country-born" by the English colonists and Scottish colonists.[98] From the second half of the 19th century to the beginning of the 20th century, new waves of immigrants from northern, eastern and southern Europe went to the Americas and consequently altered the demographics.[99] Following World War II and subsequent worldwide migrations, the current American populations' genetic admixture can be traced to all corners of the world.[35][100]
Blood groups

Prior to the 1952 confirmation of DNA as the hereditary material by Alfred Hershey and Martha Chase, scientists used blood proteins to study human genetic variation.[101][102] The ABO blood group system is widely credited to have been discovered by the Austrian Karl Landsteiner, who found three different blood types in 1900.[103] Blood groups are inherited from both parents. The ABO blood type is controlled by a single gene (the ABO gene) with three alleles: i, IA, and IB.[104]
Research by Ludwik and Hanka Herschfeld during World War I found that the frequencies of blood groups A,B and O differed greatly from region to region.[102] The "O" blood type (usually resulting from the absence of both A and B alleles) is very common around the world, with a rate of 63% in all human populations.[105] Type "O" is the primary blood type among the indigenous populations of the Americas, in-particular within Central and South America populations, with a frequency of nearly 100%.[105] In indigenous North American populations the frequency of type "A" ranges from 16% to 82%.[105] This suggests again that the initial Amerindians evolved from an isolated population with a minimal number of individuals.[106][107]
| PEOPLE GROUP | O (%) | A (%) | B (%) | AB (%) |
|---|---|---|---|---|
| Blackfoot Confederacy (N. American Indian) | 17 | 82 | 0 | 1 |
| Bororo (Brazil) | 100 | 0 | 0 | 0 |
| Eskimos (Alaska) | 38 | 44 | 13 | 5 |
| Inuit (Eastern Canada & Greenland) | 54 | 36 | 23 | 8 |
| Hawaiians | 37 | 61 | 2 | 1 |
| Indigenous North Americans (as a whole Native Nations/First Nations) | 79 | 16 | 4 | 1 |
| Maya (modern) | 98 | 1 | 1 | 1 |
| Navajo | 73 | 27 | 0 | 0 |
| Peru | 100 | 0 | 0 | 0 |
Genealogical testing
A genealogical DNA test examines the nucleotides at specific locations on a person's DNA for genetic genealogy purposes.[109] The test results are not meant to have any medical value; they are intended only to give genealogical information. Genealogical DNA tests generally involve comparing the results of living individuals to historic populations.[109] The general procedure for taking a genealogical DNA test involves taking a painless cheek-scraping (also known as a buccal swab) at home and mailing the sample to a genetic genealogy laboratory for testing. Genetic testing results showing specific sub-Haplogroups of Q, R1 and C3b implies that the individuals ancestry is, in whole or in-part, indigenous to the Americas.[110] If one's mtDNA belonged to specific sub-Haplogroups of, A, B, C, D or X2a, the implication would be that the individual's ancestry is, in whole or part, indigenous to the Americas.[111]
See also
- Archaeogenetics
- Archaeogenetics of the Near East
- Archaeology of the Americas
- Ancient DNA
- Clovis culture
- Early human migrations
- Genetic history of Africa
- Genetic history of Europe
- Genetic history of Italy
- Genetic history of North Africa
- Genetic history of the British Isles
- Genetic history of the Iberian Peninsula
- Genetics and archaeogenetics of South Asia
- Human timeline
- Life timeline
- Haplogroups of historical and famous figures
- Nature timeline
- Race and genetics
- Y-chromosome haplogroups in populations of the world
References
- ↑ Y Chromosome, Consortium (2002). "A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups". Genome Research. 12 (2): 339–348. PMC 155271
 . PMID 11827954. doi:10.1101/gr.217602.(Detailed hierarchical chart)
. PMID 11827954. doi:10.1101/gr.217602.(Detailed hierarchical chart) - 1 2 3 Griffiths, Anthony J. F. (1999). An Introduction to genetic analysis. New York: W.H. Freeman. ISBN 0-7167-3771-X. Retrieved 2010-02-03.
- 1 2 Wendy Tymchuk, Senior Technical Editor (2008). "Learn about Y-DNA Haplogroup Q. Genebase Tutorials". Genebase Systems. Archived from the original on 2010-06-22. Retrieved 2009-11-21.
- ↑ Orgel L (2004). "Prebiotic chemistry and the origin of the RNA world" (PDF). Crit Rev Biochem Mol Biol. 39 (2): 99–123. PMID 15217990. doi:10.1080/10409230490460765.
- ↑ Daniel Lee Kleinman; Kelly Moore (2014). Routledge Handbook of Science, Technology, and Society. Routledge. p. 23. ISBN 978-1-136-23716-4.
- 1 2 Tamm E, Kivisild T, Reidla M, Metspalu M, Smith DG (2007); et al. (2007), "Beringian Standstill and Spread of Native American Founders.", PLoS ONE, 2: e829, PMC 1952074
 , PMID 17786201, doi:10.1371/journal.pone.0000829
, PMID 17786201, doi:10.1371/journal.pone.0000829 - ↑ Derenko M, Malyarchuk B, Grzybowski T, Denisova G, Rogalla U (2010); et al., "Origin and Post-Glacial Dispersal of Mitochondrial DNA Haplogroups C and D in Northern Asia.", PLoS ONE, 5: e15214, PMC 3006427
 , PMID 21203537, doi:10.1371/journal.pone.0015214
, PMID 21203537, doi:10.1371/journal.pone.0015214 - ↑ Bortolini, MC; Salzano, FM; Thomas, MG; et al. (September 2003). "Y-chromosome evidence for differing ancient demographic histories in the Americas". Am. J. Hum. Genet. 73: 524–39. PMC 1180678
 . PMID 12900798. doi:10.1086/377588.
. PMID 12900798. doi:10.1086/377588. - 1 2 Ruhlen M (November 1998). "The origin of the Na-Dene". Proc. Natl. Acad. Sci. U.S.A. 95 (23): 13994–6. Bibcode:1998PNAS...9513994R. PMC 25007
 . PMID 9811914. doi:10.1073/pnas.95.23.13994.
. PMID 9811914. doi:10.1073/pnas.95.23.13994. - 1 2 Zegura SL, Karafet TM, Zhivotovsky LA, Hammer MF (January 2004). "High-resolution SNPs and microsatellite haplotypes point to a single, recent entry of Native American Y chromosomes into the Americas". Molecular Biology and Evolution. 21 (1): 164–75. PMID 14595095. doi:10.1093/molbev/msh009.
- 1 2 3 Juliette Saillard, Peter Forster, Niels Lynnerup1, Hans-Jürgen Bandelt and Søren Nørby (2000). "mtDNA Variation among Greenland Eskimos. The Edge of the Beringian Expansion". The American Journal of Human Genetics. 67 (3): 718–26. PMC 1287530
 . PMID 10924403. doi:10.1086/303038.
. PMID 10924403. doi:10.1086/303038. - ↑ Schurr, Theodore G. (2004). "The peopling of the New World – Perspectives from Molecular Anthropology". Annual Review of Anthropology. 33: 551–583. doi:10.1146/annurev.anthro.33.070203.143932.
- ↑ A. Torroni; T. G. Schurr; C. C. Yang; EJE. Szathmary; R. C. Williams; M. S. Schanfield; G. A. Troup; W. C. Knowler; D. N. Lawrence; K. M. Weiss; D. C. Wallace. "Native American Mitochondrial DNA Analysis Indicates That the Amerind and the Nadene Populations Were Founded by Two Independent Migrations". Center for Genetics and Molecular Medicine and Departments of Biochemistry and Anthropology, Emory University School of Medicine, Atlanta, Georgia. Genetics Society of America. Vol 130, 153–162. Retrieved 2009-11-28.
- ↑ "Pause Is Seen in a Continent’s Peopling". New York Times. 13 Mar 2014.
- ↑ Lyovi, Anatole (1997). An introduction to the languages of the world (Digitised online by Google books). Oxford University Press. p. 309. ISBN 0-19-508115-3. Retrieved 2010-03-25.
- ↑ Mithun, Marianne (1990). "Studies of North American Indian Languages". Annual Review of Anthropology. 19 (1): 309–330. doi:10.1146/annurev.an.19.100190.001521.
- ↑ Santos, F. R.; Pandya, A.; Tyler-Smith, C.; Pena, S. D.; Schanfield, M.; Leonard, W. R.; Mitchell, R. J. (1999). "The central Siberian origin for native American Y chromosomes". American Journal of Human Genetics. 64 (2): 619–628. PMC 1377773
 . PMID 9973301. doi:10.1086/302242.
. PMID 9973301. doi:10.1086/302242. - 1 2 Schurr, Theodore G. "Mitochondrial DNA and the Peopling of the New World" (PDF). American Scientist Online May–June 2000. Retrieved May 13, 2015.
- ↑ Zakharov, I. A., Derenko, M. V., Maliarchuk, B. A., Dambueva I. K., Dorzhu, C. M., and Rychkov, S. Y. (April 2004). "Mitochondrial DNA variation in the aboriginal populations of the Altai-Baikal region: implications for the genetic history of North Asia and America.". Ann. N. Y. Acad. Sci. 1011: 21–35. PMID 15126280. doi:10.1196/annals.1293.003.
- ↑ Starikovskaya, Elena B., Sukernik, Rem I., Derbeneva, Olga A., Volodko, Natalia A., Ruiz-Pesini, Eduardo, Torroni, Antonio, Brown, Michael D., Lott, Marie T., Hosseini, Seyed H., Huoponen, Kirsi, and Wallace, Douglas C. (January 2005). "Mitochondrial DNA diversity in indigenous populations of the southern extent of Siberia, and the origins of Native American haplogroups". Ann. Hum. Genet. 69: 67–89. PMC 3905771
 . PMID 15638829. doi:10.1046/j.1529-8817.2003.00127.x.
. PMID 15638829. doi:10.1046/j.1529-8817.2003.00127.x. - ↑ Atkinson, QD; Gray, RD; Drummond, AJ (January 2009). "Bayesian coalescent inference of major human mitochondrial DNA haplogroup expansions in Africa". Proceedings of the Royal Society B: Biological Sciences. 276 (1655): 367–73. PMC 2674340
 . PMID 18826938. doi:10.1098/rspb.2008.0785.
. PMID 18826938. doi:10.1098/rspb.2008.0785. - ↑ Watson J.D.; Crick F.H.C. (1953). "A Structure for Deoxyribose Nucleic Acid" (PDF). Nature. 171 (4356): 737–738. Bibcode:1953Natur.171..737W. PMID 13054692. doi:10.1038/171737a0. Retrieved 2009-11-27.
- ↑ "About Genetic Genealogy". Cambridge DNA Services. 2010. Retrieved 2010-01-23.
- 1 2 Ritter, Malcolm (2008). "Native American DNA Links to Six "Founding Mothers"". National Geographic Society. Retrieved 2010-01-23.
- ↑ Ayala, F (1995). "The myth of Eve:molecular biolofy and human origin". Science. 270 (5244): 1930–1936. Bibcode:1995Sci...270.1930A. PMID 8533083. doi:10.1126/science.270.5244.1930.
- ↑ Schon, Eric A. (2003). "Tales from the crypt". Department of Neurology and Department of Genetics and Development, Columbia University. American Society for Clinical Investigation. doi:10.1172/JCI20249. Retrieved 2010-01-22.
- 1 2 "After a brief review of the disciplinary context and history of physical anthropology" (PDF). Utrecht University (Universiteit Utrecht), The Netherlands. 2007. pp. 300–323. doi:10.1146/annurev.genet.41.110306.130407. Retrieved 2010-01-23.
- ↑ Underhill, Peter A.; Kivisild, Toomas (2007). "Use of Y Chromosome and Mitochondrial DNA Population Structure in Tracing Human Migrations". Annual Review of Genetics. 41: 539–564. PMID 18076332. doi:10.1146/annurev.genet.41.110306.130407.
- ↑ Silvia Fuselli; Eduardo Tarazona-Santos; Isabelle Dupanloup; Alonso Soto; Donata Luiselli; Davide Pettener (2003). "Mitochondrial DNA Diversity in South America and the Genetic History of Andean Highlanders". Mol. Biol. Evol. Society for Molecular Biology and Evolution. 20 (10): 1682–91. PMID 12832631. doi:10.1093/molbev/msg188.
- 1 2 3 4 5 Wang S, Lewis CM Jr, Jakobsson M, Ramachandran S, Ray N, et al. (2007). "Genetic Variation and Population Structure in Native Americans". PLoS Genet. 3 (11): e185. PMC 2082466
 . PMID 18039031. doi:10.1371/journal.pgen.0030185.
. PMID 18039031. doi:10.1371/journal.pgen.0030185. - ↑ Al Agellon (2008). "The Human Y Chromosomal Haplogroup Tree" (PDF). The University of Arizona. Retrieved 2010-08-27.
- 1 2 "How the Subclades of Y-DNA Haplogroup Q are determined". Genebase Systems. 2009. Retrieved 2009-11-22.
- ↑ "Y-DNA Haplogroup Tree". Genebase Systems. 2009. Retrieved 2009-11-21.
- 1 2 3 4 "Frequency Distribution of Y-DNA Haplogroup Q1a3a-M3". GeneTree. 2010. Retrieved 2010-01-30.
- 1 2 3 4 5 6 7 8 9 Wells, Spencer; Read, Mark (2002). The Journey of Man - A Genetic Odyssey (Digitised online by Google books). Random House. pp. 138–140. ISBN 0-8129-7146-9. Retrieved 2009-11-21.
- ↑ "Learning Center :: Genebase Tutorials". Genebase.com. 1964-10-22. Retrieved 2013-09-27.
- 1 2 Zegura SL, Karafet TM, Zhivotovsky LA, Hammer MF (January 2004). "High-resolution SNPs and microsatellite haplotypes point to a single, recent entry of Native American Y chromosomes into the Americas" (PDF). Mol. Biol. Evol. 21 (1): 164–75. PMID 14595095. doi:10.1093/molbev/msh009.
- ↑ Bonatto, SL; Salzano, FM (1996). "A single and early migration for the peopling of the Americas supported by mitochondrial DNA sequence data". Proceedings of the National Academy of Sciences of the United States of America. 94 (5): 1866–71. Bibcode:1997PNAS...94.1866B. PMC 20009
 . PMID 9050871. doi:10.1073/pnas.94.5.1866.
. PMID 9050871. doi:10.1073/pnas.94.5.1866. - ↑ Megan Smolenyak; Ann Turner (12 October 2004). Trace your roots with DNA: using genetic tests to explore your family tree. Rodale. pp. 83–. ISBN 978-1-59486-006-5. Retrieved 2011-06-15.
- ↑ Than, Ker (2008). "New World Settlers Took 20,000-Year Pit Stop". National Geographic Society. Retrieved 2010-01-23.
- ↑ "First Americans Arrived Recently, Settled Pacific Coast, DNA Study Says". Stefan Lovgren. National Geographic Society. 2007. Retrieved 2010-02-02.
- 1 2 3 4 Bortolini MC, Salzano FM, Thomas MG, et al. (September 2003). "Y-chromosome evidence for differing ancient demographic histories in the Americas" (PDF). Am. J. Hum. Genet. 73 (3): 524–39. PMC 1180678
 . PMID 12900798. doi:10.1086/377588.
. PMID 12900798. doi:10.1086/377588. - ↑ "First Americans Endured 20,000-Year Layover – Jennifer Viegas, Discovery News". Archived from the original on 2012-10-10. Retrieved 2009-11-18. page 2 Archived March 13, 2012, at the Wayback Machine.
- ↑ Wang, Sijia; Lewis, Cecil M.; Jakobsson, Mattias; Ramachandran, Sohini; Ray, Nicolas; Bedoya, Gabriel; Rojas, Winston; Parra, Maria V.; Molina, Julio A. (2007). "Genetic Variation and Population Structure in Native Americans". PLoS Genetics. 3 (11): e185. PMC 2082466
 . PMID 18039031. doi:10.1371/journal.pgen.0030185.
. PMID 18039031. doi:10.1371/journal.pgen.0030185. - ↑ Malhi (2008). "Distribution of Y Chromosomes Among Native North Americans: A Study of Athapaskan Population History". American Journal of Physical Anthropology. 137: 412–24. PMC 2584155
 . PMID 18618732. doi:10.1002/ajpa.20883.
. PMID 18618732. doi:10.1002/ajpa.20883. - ↑ González Burchard, E; Borrell, LN; Choudhry, S; Naqvi, M; Tsai, HJ; Rodriguez-Santana, JR; Chapela, R; Rogers, SD; Mei, R (December 2005). "Latino Populations: A Unique Opportunity for the Study of Race, Genetics, and Social Environment in Epidemiological Research". American Journal of Public Health. 95 (12): 2161–8. PMC 1449501
 . PMID 16257940. doi:10.2105/AJPH.2005.068668.
. PMID 16257940. doi:10.2105/AJPH.2005.068668. - 1 2 3 Bortolini, MC; Salzano, FM; Thomas, MG; et al. (2003). "Y-chromosome evidence for differing ancient demographic histories in the Americas". Am. J. Hum. Genet. 73 (3): 524–39. PMC 1180678
 . PMID 12900798. doi:10.1086/377588.
. PMID 12900798. doi:10.1086/377588. - 1 2 3 Singh, Ripan (2008). "Distribution of Y Chromosomes Among Native North Americans: A Study of Athapaskan Population History" (PDF). American Journal of Physical Anthropology. Retrieved 2010-08-27.
- ↑ Bortolini, MC; Salzano, FM; Thomas, MG; et al. "(September 2003) "Y-chromosome evidence for differing ancient demographic histories in the Americas". Am. J. Hum. Genet. 73 (3): 524–39. PMC 1180678
 . PMID 12900798. doi:10.1086/377588.
. PMID 12900798. doi:10.1086/377588. - ↑ Lell, Jeffrey T.; Sukernik, Rem I.; Starikovskaya, Yelena B.; Su, Bing; Jin, Li; Schurr, Theodore G.; Underhill, Peter A.; Wallace, Douglas C. (2002). "The Dual Origin and Siberian Affinities of Native American". The American Journal of Human Genetics. 70 (1): 192–206. PMC 384887
 . PMID 11731934. doi:10.1086/338457.
. PMID 11731934. doi:10.1086/338457. - 1 2 3 Raghavan, Maanasa; Skoglund, Pontus; Graf, Kelly E.; Metspalu, Mait; Albrechtsen, Anders; Moltke, Ida; Rasmussen, Simon; Thomas, W. Stafford Jr; Orlando, Ludovic; Metspalu, Ene; Karmin, Monika; Tambets, Kristiina; Rootsi, Siiri; Mägi, Reedik; Campos, Paula F.; Balanovska, Elena; Balanovsky, Oleg; Khusnutdinova, Elza; Litvinov, Sergey; Osipova, Ludmila P.; Fedorova, Sardana A.; Voevoda, Mikhail I.; DeGiorgio, Michael; Sicheritz-Ponten, Thomas; Brunak, Søren; et al. (2014). "Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans". Nature. 505 (7481): 87–91. PMC 4105016
 . PMID 24256729. doi:10.1038/nature12736.
. PMID 24256729. doi:10.1038/nature12736. - ↑ Michael Balter (October 2013). "Ancient DNA Links Native Americans With Europe". Science. 342 (6157): 409–410. Bibcode:2013Sci...342..409B. PMID 24159019. doi:10.1126/science.342.6157.409.
- ↑ Xue Y, Zerjal T, Bao W, et al. (April 2006). "Male demography in East Asia: a north-south contrast in human population expansion times". Genetics. 172 (4): 2431–9. PMC 1456369
 . PMID 16489223. doi:10.1534/genetics.105.054270.
. PMID 16489223. doi:10.1534/genetics.105.054270. - ↑ BRUCE JOHANSEN; BARRY PRITZKER (23 July 2007). Encyclopedia of American Indian History. ABC-CLIO. p. 441. ISBN 978-1-85109-818-7.
- ↑ Cann RL, Stoneking M, Wilson AC (1987), "Mitochondrial DNA and human evolution", Nature, 325 (6099): 31–36, Bibcode:1987Natur.325...31C, PMID 3025745, doi:10.1038/325031a0
- ↑ Soares P, Ermini L, Thomson N, et al. (June 2009), "Correcting for purifying selection: an improved human mitochondrial molecular clock", Am. J. Hum. Genet., 84 (6): 740–59, PMC 2694979
 , PMID 19500773, doi:10.1016/j.ajhg.2009.05.001.
University of Leeds – New 'molecular clock' aids dating of human migration history
, PMID 19500773, doi:10.1016/j.ajhg.2009.05.001.
University of Leeds – New 'molecular clock' aids dating of human migration history - ↑ Takahata, N (January 1993). "Allelic genealogy and human evolution". Mol. Biol. Evol. 10 (1): 2–22. PMID 8450756.
- ↑ Dawkins, Richard (2004). The ancestor's tale: a pilgrimage to the dawn of evolution. Boston: Houghton Mifflin. p. 64. ISBN 0-618-00583-8. Retrieved 2010-01-24.
- 1 2 3 Jason A. Eshleman; Ripan S. Malhi; David Gleen Smith (2003). "Mitochondrial DNA Studies of Native Americans- Conceptions and Misconceptions of the Population Prehistory of the Americas" (PDF). 7. Retrieved 2010-01-23.
- 1 2 3 Achilli A, Perego UA, Bravi CM, Coble MD, Kong Q-P; Perego; Bravi; Coble; Kong; Woodward; Salas; Torroni; Bandelt (2008). MacAulay, Vincent, ed. "The Phylogeny of the Four Pan-American MtDNA Haplogroups: Implications for Evolutionary and Disease Studies". PLoS ONE. 3 (3): e1764. Bibcode:2008PLoSO...3.1764A. PMC 2258150
 . PMID 18335039. doi:10.1371/journal.pone.0001764.
. PMID 18335039. doi:10.1371/journal.pone.0001764. - ↑ "Learn about Y-DNA Haplogroup Q". Wendy Tymchuk, Senior Technical Editor. Genebase Systems. 2008. Archived from the original on 2010-06-22. Retrieved 2009-11-21.
- ↑ "The peopling of the Americas: Genetic ancestry influences health". Scientific American. Retrieved 2009-11-27.
- 1 2 Fagundes, Nelson J.R.; Ricardo Kanitz, Roberta Eckert, Ana C.S. Valls, Mauricio R. Bogo, Francisco M. Salzano, David Glenn Smith, Wilson A. Silva, Marco A. Zago, Andrea K. Ribeiro-dos-Santos, Sidney E.B. Santos, Maria Luiza Petzl-Erler, and Sandro L.Bonatto (2008). "Mitochondrial Population Genomics Supports a Single Pre-Clovis Origin with a Coastal Route for the Peopling of the Americas" (PDF). American Journal of Human Genetics. 82 (3): 583–592. PMC 2427228
 . PMID 18313026. doi:10.1016/j.ajhg.2007.11.013.
. PMID 18313026. doi:10.1016/j.ajhg.2007.11.013. - ↑ David J. Meltzer (2009). First Peoples in a New World: Colonizing Ice Age America. University of California Press. p. 162. ISBN 978-0-520-94315-5.
- ↑ "An mtDNA view of the peopling of the world by Homo sapiens". Cambridge DNA. 2009. Archived from the original on 2011-05-11. Retrieved 2010-04-29.
- ↑ Reidla, M; Kivisild, T; Metspalu, E; Kaldma, K; Tambets, K; Tolk, HV; Parik, J; Loogväli, EL; et al. (2003). "Origin and Diffusion of mtDNA Haplogroup X". Am J Hum Genet. 73 (5): 1178–90. PMC 1180497
 . PMID 14574647. doi:10.1086/379380.
. PMID 14574647. doi:10.1086/379380. - ↑ "An mtDNA view of the peopling of the world by Homo sapiens". Cambridge DNA Services. 2007. Archived from the original on 2011-05-11. Retrieved 2011-06-01.
- 1 2 Robert E. Ferrell; Ranajit Chakraborty; Henry Gershowitz; W. S. Laughlin; W. J. Schull (1990). "The St. Lawrence Island Eskimos, Genetic Variation and Genetic Distance". American Journal of Physical Anthropology. 55 (3): 351–8. PMID 6455922. doi:10.1002/ajpa.1330550309.
- 1 2 Helgason, Agnar; Pálsson, Gísli; Pedersen, Henning Sloth; Angulalik, Emily; Gunnarsdóttir, Ellen Dröfn; Yngvadóttir, Bryndís; Stefánsson, Kári (2006). "mtDNA variation in Inuit populations of Greenland and Canada: migration history". American Journal of Physical Anthropology. 130 (1): 123–134. PMID 16353217. doi:10.1002/ajpa.20313.
- ↑ Raghavan, M; Skoglund, P; Graf, KE; et al. (January 2014). "Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans". Nature. 505: 87–91. PMC 4105016
 . PMID 24256729. doi:10.1038/nature12736. Lay summary – ScienceDaily.
. PMID 24256729. doi:10.1038/nature12736. Lay summary – ScienceDaily. - ↑ "Ancient Siberian Genome Reveals Genetic Origins of Native Americans". PHYSORG. Nov 20, 2013. Retrieved 2013-11-23.
- ↑ Kashani, Baharak Hooshiar; Ugo A. Perego1, Anna Olivieri, Norman Angerhofer, Francesca Gandini, Valeria Carossa, Hovirag Lancioni, Ornella Semino, Scott R. Woodward, Alessandro Achilli, Antonio Torroni (January 2012). "Mitochondrial haplogroup C4c: A rare lineage entering America through the ice-free corridor?". American Journal of Physical Anthropology. 147 (1): 35–39. PMID 22024980. doi:10.1002/ajpa.21614.
- ↑ "PLOS ONE". plosone.org. Retrieved 10 November 2015.
- ↑ Ancient mitochondrial DNA provides high-resolution time scale of the peopling of the Americas, 2016
- 1 2 3 4 "Joint match probabilities for Y chromosomal and autosomal markers" (PDF). Forensic Science International. 2008. pp. 174, 234–238. Archived from the original (PDF) on 2010-07-03. Retrieved 2010-02-03.
- 1 2 Hey, J (2005). "On the Number of New World Founders: A Population Genetic Portrait of the Peopling of the Americas". Hey J: PLoS Biol. 3: e193. PMC 1131883
 . PMID 15898833. doi:10.1371/journal.pbio.0030193. Retrieved 2014-03-24.
. PMID 15898833. doi:10.1371/journal.pbio.0030193. Retrieved 2014-03-24. - ↑ Wade, Nicholas (2010). "Ancient Man In Greenland Has Genome Decoded". The New York Times Company. Retrieved 2010-01-10.
- ↑ Rasmussen, Morten; Anzick, Sarah L.; Waters, Michael R.; Skoglund, Pontus; Degiorgio, Michael; Stafford, Thomas W.; Rasmussen, Simon; Moltke, Ida; Albrechtsen, Anders; Doyle, Shane M.; Poznik, G. David; Gudmundsdottir, Valborg; Yadav, Rachita; Malaspinas, Anna-Sapfo; v, Samuel Stockton White; Allentoft, Morten E.; Cornejo, Omar E.; Tambets, Kristiina; Eriksson, Anders; Heintzman, Peter D.; Karmin, Monika; Korneliussen, Thorfinn Sand; Meltzer, David J.; Pierre, Tracey L.; Stenderup, Jesper; Saag, Lauri; Warmuth, Vera M.; Lopes, Margarida C.; Malhi, Ripan S.; et al. (2014). "The genome of a Late Pleistocene human from a Clovis burial site in western Montana". Nature. 506 (7487): 225–229. Bibcode:2014Natur.506..225R. PMID 24522598. doi:10.1038/nature13025.
- ↑ "Ancient American's genome mapped". BBC News. 2014-02-14.
- 1 2 "Reconstructing Native American population history". Nature. 488: 370–374. doi:10.1038/nature11258. Retrieved 10 November 2015.
- ↑ Abstract Profiles of Structural Stability Point to Universal Tendencies, Family-Specific Factors, and Ancient Connections between Languages, 2012
- ↑ Raghavan; et al. (21 August 2015). "Genomic evidence for the Pleistocene and recent population history of Native Americans" (pdf). Retrieved 6 October 2015.
- ↑ Skoglund, P; Mallick, S; Bortolini, MC; Chennagiri, N; Hünemeier, T; Petzl-Erler, ML; Salzano, FM; Patterson, N; Reich, D (21 July 2015). "Genetic evidence for two founding populations of the Americas" (pdf). Nature. 525: 104–8. PMC 4982469
 . PMID 26196601. doi:10.1038/nature14895. Retrieved 7 October 2015.
. PMID 26196601. doi:10.1038/nature14895. Retrieved 7 October 2015. - 1 2 Kimura M (June 1962). "On the probability of fixation of mutant genes in a population". Genetics. 47 (6): 713–9. PMC 1210364
 . PMID 14456043. Retrieved 2010-02-03.
. PMID 14456043. Retrieved 2010-02-03. - ↑ "The peopling of the Americas: Genetic ancestry influences health". scientific journal American Journal of Physical Anthropology. University of Oklahoma. 2009. Retrieved 2009-11-21.
- ↑ Díaz-Matallana, Marcela (2009). "To the Caribbean: Indigenous MtDNA" (PDF). University of Puerto Rico. Molecular Population Genetics. Retrieved 2010-01-22.
- ↑ Weber, George (2007). Fuegian and Patagonian Genetics and the settling of the Americas. Andaman Association. Retrieved 2010-01-22.
García-Bour J, Pérez-Pérez A, Alvarez S, et al. (April 2004). "Early population differentiation in extinct aborigines from Tierra del Fuego-Patagonia: ancient mtDNA sequences and Y-chromosome STR characterization". Am. J. Phys. Anthropol. 123 (4): 361–70. PMID 15022364. doi:10.1002/ajpa.10337. - ↑ Bjerregaard P, Jørgensen ME, Borch-Johnsen K (June 2004). "Serum lipids of Greenland Inuit in relation to Inuit genetic heritage, westernisation and migration". Atherosclerosis. 174 (2): 391–8. PMID 15136072. doi:10.1016/j.atherosclerosis.2004.02.010.
- ↑ Leffell MS, Fallin MD, Erlich HA, et al. (July 2002). "HLA antigens, alleles and haplotypes among the Yup'ik Alaska natives: report of the ASHI Minority Workshops, Part II". Human Immunology. 63 (7): 614–25. PMID 12072196. doi:10.1016/S0198-8859(02)00415-9.
- ↑ Newman, Richard (1999). "Miscegenation". In Kwame Appiah and Henry Louis Gates, Jr. Africana: The Encyclopedia of the African and African American Experience (1st ed.). New York: Basic Civitas Books. p. 1320. ISBN 0-465-00071-1.
Miscegenation, a term for sexual relations across racial lines; no longer in use because of its racist implications
- ↑ Arnold, David (2002). The Age of Discovery, 1400–1600. Lancaster pamphlets, Routledge. p. 11. ISBN 0-415-27996-8.
- 1 2 3 Chasteen, John Charles; Wood, James A (2003). Problems in modern Latin American history, sources and interpretations (Digitized online by Google books). Sr Books. pp. 4–10. ISBN 0-8420-5060-4. Retrieved 2010-02-24.
- ↑ "Admixture Studies in Latin America: From the 20th to the 21st Century" (PDF). Facultad de Humanidades y Ciencias de la Educación. 2000. Retrieved 2010-02-24.
- ↑ Sweet, Frank W (2004). "Afro-European Genetic Admixture in the United States". Essays on the Color Line and the One-Drop Rule. Backintyme Essays. Retrieved 2010-02-24.
- ↑ Sweet, Frank W (2005). Legal History of the Color Line: The Notion of Invisible Blackness. Backintyme Publishing. p. 542. ISBN 0-939479-23-0. Retrieved 2010-02-24.
- ↑ Soong, Roland (1999). "Racial Classifications in Latin America". Latin American Media. Retrieved 2010-02-25.
- ↑ Brehaut, Harry B (1998). "The Red River Cart and Trails: The Fur Trade". Manitoba Historical Society. Retrieved 2010-02-24.
- ↑ "Who are Métis ?". Métis National Council. 2001. Retrieved 2010-02-24.
- ↑ Wagenknecht, Malte (2004). International migration during the 19th century (Digitized online by Google books). Åbo Akademi University. ISBN 978-3-638-78306-4. Retrieved 2010-02-24.
- ↑ "Genetic Anthropology, Ancestry, and Ancient Human Migration". The U.S. Department of Energy Office of Science, Office of Biological and Environmental Research, Human Genome Program. 2010. Retrieved 2014-03-24.
- ↑ Hershey A, Chase M (1952). "Independent functions of viral protein and nucleic acid in growth of bacteriophage" (PDF). J Gen Physiol. 36 (1): 39–56. PMC 2147348
 . PMID 12981234. doi:10.1085/jgp.36.1.39.
. PMID 12981234. doi:10.1085/jgp.36.1.39. - 1 2 LILLE-SZYSZKOWICZ I (1957). "Development of studies on pleiades of blood groups". Postȩpy higieny i medycyny doświadczalnej. 11 (3): 229–33. PMID 13505351.
- ↑ Landsteiner K (1900). "Zur Kenntnis der antifermentativen, lytischen und agglutinierenden Wirkungen des Blutserums und der Lymphe". Zentralblatt Bakteriologie. 27: 357–62.
- ↑ Yazer M, Olsson M, Palcic M (2006). "The cis-AB blood group phenotype: fundamental lessons in glycobiology". Transfus Med Rev. 20 (3): 207–17. PMID 16787828. doi:10.1016/j.tmrv.2006.03.002.
- 1 2 3 "Distribution of Blood Types". Behavioral Sciences Department, Palomar College. 2010. Archived from the original on 2006-02-21. Retrieved 2010-03-14.
- ↑ Estrada-Mena B, Estrada FJ, Ulloa-Arvizu R, et al. (May 2010). "Blood group O alleles in Native Americans: implications in the peopling of the Americas". Am. J. Phys. Anthropol. 142 (1): 85–94. PMID 19862808. doi:10.1002/ajpa.21204.
- ↑ Chown, B; Lewis, M (2005). "The blood group genes of the Cree Indians and the Eskimos of the ungava district of Canada". American Journal of Physical Anthropology. 14 (2): 215–224. PMID 13362488. doi:10.1002/ajpa.1330140217.
- ↑ "Racial and Ethnic Distribution of ABO Blood Types". BloodBook. 2008. Retrieved 2010-03-31.
- 1 2 Anne Hart; Anne Hart M a (April 2004). How to Interpret Family History and Ancestry DNA Test Results for Beginners: The Geography and History of Your Relatives. iUniverse. pp. 449–. ISBN 978-0-595-31684-7. Retrieved 2011-06-15.
- ↑ Blanco Verea; Alejandro José. Linajes del cromosoma Y humano: aplicaciones genético-poblacionales y forenses. Univ Santiago de Compostela. pp. 135–. GGKEY:JCW0ASCR364. Retrieved 2011-06-15.
- ↑ Nina G. Jablonski (2002). The first Americans: the Pleistocene colonization of the New World. University of California Press. p. 301. ISBN 978-0-940228-50-4. Retrieved 2011-06-15.
Further reading
- Peter N. Jones (October 2002). American Indian mtDNA, Y chromosome genetic data, and the peopling of North America. Bauu Institute. ISBN 978-0-9721349-1-0.
- Joseph Frederick Powell (2005). The first Americans: race, evolution, and the origin of Native Americans. Cambridge University Press. ISBN 978-0-521-82350-0.
- Francisco M. Salzano; Maria Cátira Bortolini (2002). The evolution and genetics of Latin American populations. Cambridge University Press. ISBN 978-0-521-65275-9.
External links
| Wikimedia Commons has media related to Genetic studies of indigenous populations of the Americas. |
- The Great Human Odyssey - Canadian Broadcasting Corporation
- Atlas of the Human Journey, Genographic Project, National Geographic
- Journey of Mankind – Genetic Map – Bradshaw Foundation
- World Haplogroups Maps (2005) – University of Illinois
- Q yDNA Project – International society of genetic genealogy
- Human Timeline (Interactive) – Smithsonian, National Museum of Natural History (August 2016).
.svg.png)