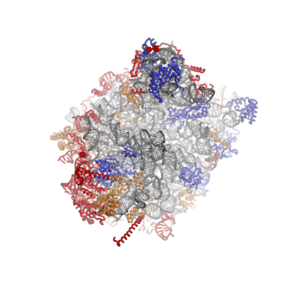
Eukaryotic large ribosomal subunit (60S)
Ribosomal particles are denoted according to their sedimentation coefficients in Svedberg units. The 60S subunit is the large subunit of eukaryotic 80S ribosomes. It is structurally and functionally related to the 50S subunit of 70S prokaryotic ribosomes.[1][2][3][4][5][6] However, the 60S subunit is much larger than the prokaryotic 50S subunit and contains many additional protein segments, as well as ribosomal RNA expansion segments.
Overall structure
Characteristic features of the large subunit, shown below in the "Crown View", include the central protuberance (CP) and the two stalks, which are named according to their bacterial protein components (L1 stalk on the left as seen from the subunit interface and L7/L12 on the right). There are three binding sites for tRNA, the A-site, P-site and E-site (see article on protein translation for details). The core of the 60S subunit is formed by the 28S ribosomal RNA (abbreviated 28S rRNA), which is homologous to the prokaryotic 23S rRNA, which also contributes the active site (peptidyl transferase center, PTC) of the ribosome.[2][4] The rRNA core is decorated with dozens of proteins. In the figure "Crystal Structure of the Eukaryotic 60S Ribosomal Subunit from T. thermophila", the ribosomal RNA core is represented as a grey tube and expansion segments are shown in red. Proteins which have homologs in eukaryotes, archaea and bacteria are shown as blue ribbons. Proteins shared only between eukaryotes and archaea are shown as orange ribbons and proteins specific to eukaryotes are shown as red ribbons.
| Crystal Structure of the Eukaryotic 60S Ribosomal Subunit from T. thermophila | ||||
|---|---|---|---|---|
|
60S ribosomal proteins
The table "60S ribosomal proteins" shows the individual protein folds of the 60S subunit colored by conservation as above. The eukaryote-specific extensions, ranging from a few residues or loops to very long alpha helices and additional domains, are highlighted in red.[2]
Historically, different nomenclatures have been used for ribosomal proteins. For instance, proteins have been numbered according to their migration properties in gel electrophoresis experiments. Therefore, different names may refer to homologous proteins from different organisms, while identical names do not necessarily denote homologous proteins. The table "60S ribosomal proteins" cross-references the human ribosomal protein names with yeast, bacterial, and archaeal homologs.[7] Further information can be found in the ribosomal protein gene database (RPG).[7]
| Structure (Eukaryotic)[8] | H. sapiens[7][9] | Amino acids[10] | Conservation[11] | S. cerevisiae[12] | Bacterial homolog (E. coli) | Archaeal homolog |
|---|---|---|---|---|---|---|
 | RPLP0 | 318 | EAB | P0 | L10 | L10 |
 | RPL3 | 404 | EAB | L3 | L3 | L3 |
 | RPL4 | 428 | EAB | L4 | L4 | L4 |
 | RPL5 | 298 | EAB | L5 | L18 | L18p |
 | RPL6 | 289 | E | L6 | n/a | n/a |
 | RPL7 | 254 | EAB | L7 | L30 | L30 |
 | RPL7A | 267 | EA | L8 | n/a | L7Ae |
 | RPL8 | 258 | EAB | L2 | L2 | L2 |
 | RPL9 | 193 | EAB | L9 | L6 | L6 |
 | RPL10 | 215 | EAB | L10 | L16 | L10e |
 | RPL11 | EAB | L11 | L5 | L5 | |
 | RPL13 | EA | L13 | n/a | L13e | |
 | RPL13A | 204 | EAB | L16 | L13 | L13 |
 | RPL14 | 221 | EA | L14 | n/a | L14e |
 | RPL15 | 205 | EA | L15 | n/a | L15e |
 | RPL17 | 185 | EAB | L17 | L22 | L22 |
 | RPL18 | 189 | EA | L18 | n/a | L18e |
 | RPL18A | 177 | EA | L20 | n/a | Lx |
 | RPL19 | 197 | EA | L19 | n/a | L19 |
 | RPL21 | 161 | EA | L21 | n/a | L21e |
 | RPL22 | 129 | E | L22 | n/a | n/a |
 | RPL23 | 141 | EAB | L23 | L14 | L14p |
 | RPL23A | 157 | EAB | L25 | L23 | L23 |
 | RPL24 | 158 | EA | L24 | n/a | L24e |
 | RPL26 | 146 | EAB | L26 | L24 | L24 |
 | RPL27 | 137 | E | L27 | n/a | n/a |
 | RPL27A | 149 | EAB | L28 | L15 | L15 |
 | RPL28 | E | n/a[2][3][13] | n/a | n/a | |
 | RPL29 | E | L29 | n/a | n/a | |
 | RPL30 | 116 | EA | L30 | n/a | L30e |
 | RPL31 | 126 | EA | L31 | n/a | L31e |
 | RPL32 | 136 | EA | L32 | n/a | L32e |
 | RPL34 | 118 | EA | L34 | n/a | L34e |
 | RPL35 | 124 | EAB | L35 | L29 | L29 |
 | RPL35A | EA | L33 | n/a | L35Ae | |
 | RPL36 | 106 | E | L36 | n/a | n/a |
 | RPL36A | 107 | EA | L42 | n/a | L44e |
 | RPL37 | 98 | EA | L37 | n/a | L37e |
 | RPL37A | EA | L43 | n/a | L37Ae | |
 | RPL38 | EA | L38 | n/a | L38e | |
 | RPL39 | 52 | EA | L39 | n/a | L37Ae |
 | RPL40 | 129 | EA | L40 | n/a | L40e |
References
- ↑ 60S Ribosome Subunits at the US National Library of Medicine Medical Subject Headings (MeSH)
- 1 2 3 4 Klinge S, Voigts-Hoffmann F, Leibundgut M, Arpagaus S, Ban N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science. 2011 Nov 18;334(6058):941-8. doi:10.1126/science.1211204. Epub 2011 Nov 3. PMID 22052974.
- 1 2 Ben-Shem A, Garreau de Loubresse N, Melnikov S, Jenner L, Yusupova G, Yusupov M. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science. 2011 Dec 16;334(6062):1524-9. doi:10.1126/science.1212642. Epub 2011 Nov 17. PMID 22096102.
- 1 2 Ban N, Nissen P, Hansen J, Moore PB, Steitz TA. The complete atomic structure of the large ribosomal subunit at 2.4 A resolution. Science. 2000 Aug 11;289(5481):905-20. PMID 10937989.
- ↑ Cate JH, Yusupov MM, Yusupova GZ, Earnest TN, Noller HF. X-ray crystal structures of 70S ribosome functional complexes. Science. 1999 Sep 24;285(5436):2095-104. PMID 10497122.
- ↑ Yusupov MM, Yusupova GZ, Baucom A, Lieberman K, Earnest TN, Cate JH, Noller HF. Crystal structure of the ribosome at 5.5 A resolution. Science. 2001 May 4;292(5518):883-96. Epub 2001 Mar 29. PMID 11283358.
- 1 2 3 Nakao A, Yoshihama M, Kenmochi N. RPG: the Ribosomal Protein Gene database. Nucleic Acids Res. 2004 Jan 1;32(Database issue):D168-70. PMID 14681386; PMC 308739.
- ↑ Structure of the T. thermophila,' proteins from the structures of the large subunit PDBS 417, 4A19
- ↑ Nomenclature according to the ribosomal protein gene database, applies to H. sapiens and T. thermophila
- ↑ Yoshihama, Maki; Uechi, Tamayo; Asakawa, Shuichi; Kawasaki, Kauhiko (2002). "The Human Ribosomal Protein Genes: Sequencing and Comparative Analysis of 73 Genes". Genome Research. 12: 379–390. PMC 155282
 . doi:10.1101/gr.214202.
. doi:10.1101/gr.214202. - ↑ EAB means conserved in eukaryotes, archaea and bacteria, EA means conserved in eukaryotes and archaea and E means eukaryote-specific protein
- ↑ Traditionally, ribosomal proteins were named according to their apparent molecular weight in gel electrophoresis, leading to different names for homologous proteins from different organisms. The RPG offers a unified nomenclature for ribosomal protein genes based on homology.
- ↑ RPL28 has no detectable homolog in yeast