dUTP diphosphatase
| dUTP diphosphatase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC number | 3.6.1.23 | ||||||||
| CAS number | 37289-34-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
| dUTPase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
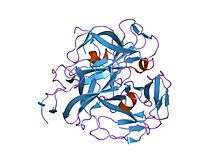 crystal structures of feline immunodeficiency virus dutp pyrophosphatase and its nucleotide complexes in three crystal forms. | |||||||||
| Identifiers | |||||||||
| Symbol | dUTPase | ||||||||
| Pfam | PF00692 | ||||||||
| Pfam clan | CL0153 | ||||||||
| InterPro | IPR008180 | ||||||||
| SCOP | 1dup | ||||||||
| SUPERFAMILY | 1dup | ||||||||
| |||||||||
| dUTPase_2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 the crystal structure of a complex of campylobacter jejuni dutpase with substrate analogue dupnhp | |||||||||
| Identifiers | |||||||||
| Symbol | dUTPase_2 | ||||||||
| Pfam | PF08761 | ||||||||
| Pfam clan | CL0231 | ||||||||
| InterPro | IPR014871 | ||||||||
| SCOP | 1w2y | ||||||||
| SUPERFAMILY | 1w2y | ||||||||
| |||||||||
In enzymology, a dUTP diphosphatase (EC 3.6.1.23) is an enzyme that catalyzes the chemical reaction
- dUTP + H2O dUMP + diphosphate
Thus, the two substrates of this enzyme are dUTP and H2O, whereas its two products are dUMP and diphosphate.
This enzyme belongs to the family of hydrolases, specifically those acting on acid anhydrides in phosphorus-containing anhydrides. The systematic name of this enzyme class is dUTP nucleotidohydrolase. Other names in common use include deoxyuridine-triphosphatase, dUTPase, dUTP pyrophosphatase, desoxyuridine 5'-triphosphate nucleotidohydrolase, and desoxyuridine 5'-triphosphatase. This enzyme participates in pyrimidine metabolism.
This enzyme has a dual function: on one hand, it removes dUTP from the deoxynucleotide pool, which reduces the probability of this base being incorporated into DNA by DNA polymerases, while on the other hand, it produces the dTTP precursor dUMP. Lack or inhibition of dUTPase action leads to harmful perturbations in the nucleotide pool resulting in increased uracil content of DNA that activates a hyperactive futile cycle of DNA repair.[1][2]
Structural studies
As of late 2007, 48 structures have been solved for this class of enzymes, with PDB accession codes 1DUC, 1DUD, 1DUN, 1DUP, 1DUT, 1EU5, 1EUW, 1F7D, 1F7K, 1F7N, 1F7O, 1F7P, 1F7Q, 1F7R, 1MQ7, 1OGH, 1OGK, 1OGL, 1PKH, 1PKJ, 1PKK, 1RN8, 1RNJ, 1SEH, 1SIX, 1SJN, 1SLH, 1SM8, 1SMC, 1SNF, 1SYL, 1VYQ, 1W2Y, 2BSY, 2BT1, 2CJE, 2D4L, 2D4M, 2D4N, 2HQU, 2HR6, 2HRM, 2OKB, 2OKD, 2OKE, 2OL0, 2OL1, and 2PY4.
There are at least two structurally distinct families of dUTPases. The crystal structure of human dUTPase reveals that each subunit of the dUTPase trimer folds into an eight-stranded jelly-roll beta barrel, with the C-terminal beta strands interchanged among the subunits. The structure is similar to that of the Escherichia coli enzyme, despite low sequence homology between the two enzymes.[3]
The second family has a novel all-alpha fold, members of this family are unrelated to the all-beta fold found in dUTPases of the majority of organisms.[4]
References
- ↑ Vertessy BG, Toth J (2009). "Keeping uracil out of DNA". Accounts of Chemical Research. 42 (1): 97–106. PMC 2732909
 . PMID 18837522. doi:10.1021/ar800114w.
. PMID 18837522. doi:10.1021/ar800114w. - ↑ Vassylyev DG, Morikawa K (1996). "Precluding uracil from DNA". Structure. 4 (12): 1381–5. PMID 8994964. doi:10.1016/S0969-2126(96)00145-1.
- ↑ Mol CD, Harris JM, McIntosh EM, Tainer JA (September 1996). "Human dUTP pyrophosphatase: uracil recognition by a beta hairpin and active sites formed by three separate subunits". Structure. 4 (9): 1077–92. PMID 8805593. doi:10.1016/S0969-2126(96)00114-1.
- ↑ Moroz, O. V.; Harkiolaki, M.; Galperin, M. Y.; Vagin, A. A.; González-Pacanowska, D.; Wilson, K. S. (2004). "The Crystal Structure of a Complex of Campylobacter jejuni dUTPase with Substrate Analogue Sheds Light on the Mechanism and Suggests the "Basic Module" for Dimeric d(C/U)TPases". Journal of Molecular Biology. 342 (5): 1583–1597. PMID 15364583. doi:10.1016/j.jmb.2004.07.050.
Further reading
- Bertani Le.; Haeggmark A.; Reichard P.; Interconversion of Deoxyuridine Phosphates (1963). "Enzymatic Synthesis of Deoxyribonucleotides. II. Formation". J. Biol. Chem. 238: 3407–13. PMID 14085395.
- Giroir LE, Deutsch WA (1987). "Drosophila deoxyuridine triphosphatase. Purification and characterization". J. Biol. Chem. 262 (1): 130–4. PMID 3025197.
- Greenberg G, Somerville R (1962). "DEOXYURIDYLATE KINASE ACTIVITY AND DEOXYURIDINETRIPHOSPHATASE IN ESCHERICHIA COLI". Proc. Natl. Acad. Sci. U.S.A. 48 (2): 247–57. PMC 220766
 . PMID 13901467. doi:10.1073/pnas.48.2.247.
. PMID 13901467. doi:10.1073/pnas.48.2.247. - Grindey GR, Nichol CA (1971). "Mammalian deoxyuridine 5'-triphosphate pyrophosphatase". Biochim. Biophys. Acta. 240 (2): 180–3. PMID 5105331. doi:10.1016/0005-2787(71)90655-1.
This article incorporates text from the public domain Pfam and InterPro IPR008180
This article incorporates text from the public domain Pfam and InterPro IPR014871