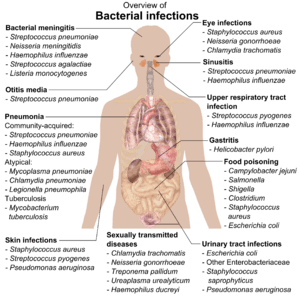
Bacteria
| Bacteria Temporal range: Archean or earlier – present | |
|---|---|
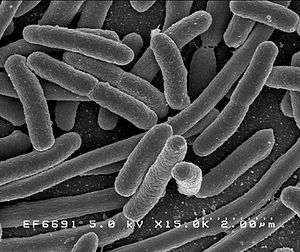 | |
| Scanning electron micrograph of Escherichia coli rods | |
| Scientific classification | |
| Domain: | Bacteria Woese, Kandler & Wheelis, 1990[1] |
| Phyla | |
|
Actinobacteria (high-G+C)
Aquificae
Acidobacteria | |
| Synonyms | |
|
Eubacteria Woese & Fox, 1977[2] | |
Bacteria (/bækˈtɪəriə/; common noun bacteria, singular bacterium) constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria have a number of shapes, ranging from spheres to rods and spirals. Bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste,[3] and the deep portions of Earth's crust. Bacteria also live in symbiotic and parasitic relationships with plants and animals. Most bacteria have not been characterised, and only about half of the bacterial phyla have species that can be grown in the laboratory.[4] The study of bacteria is known as bacteriology, a branch of microbiology.
There are typically 40 million bacterial cells in a gram of soil and a million bacterial cells in a millilitre of fresh water. There are approximately 5×1030 bacteria on Earth,[5] forming a biomass which exceeds that of all plants and animals.[6] Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies and bacteria are responsible for the putrefaction stage in this process.[7] In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. In March 2013, data reported by researchers in October 2012, was published. It was suggested that bacteria thrive in the Mariana Trench, which with a depth of up to 11 kilometres is the deepest known part of the oceans.[8][9] Other researchers reported related studies that microbes thrive inside rocks up to 580 metres below the sea floor under 2.6 kilometres of ocean off the coast of the northwestern United States.[8][10] According to one of the researchers, "You can find microbes everywhere—they're extremely adaptable to conditions, and survive wherever they are."[8]
The famous notion that bacterial cells outnumber human cells by a factor of 10:1 has been debunked. There are approximately 39 trillion bacterial cells in the human microbiota as personified by an "reference" 70 kg male 170 cm tall, whereas there are 30 trillion human cells in the body. This means that although they do have the upper hand in actual numbers, it is only by 30%, and not 900%.[11]
The largest number exist in the gut flora, and a large number on the skin.[12] The vast majority of the bacteria in the body are rendered harmless by the protective effects of the immune system, though many are beneficial particularly in the gut flora. However several species of bacteria are pathogenic and cause infectious diseases, including cholera, syphilis, anthrax, leprosy, and bubonic plague. The most common fatal bacterial diseases are respiratory infections, with tuberculosis alone killing about 2 million people per year, mostly in sub-Saharan Africa.[13] In developed countries, antibiotics are used to treat bacterial infections and are also used in farming, making antibiotic resistance a growing problem. In industry, bacteria are important in sewage treatment and the breakdown of oil spills, the production of cheese and yogurt through fermentation, and the recovery of gold, palladium, copper and other metals in the mining sector,[14] as well as in biotechnology, and the manufacture of antibiotics and other chemicals.[15]
Once regarded as plants constituting the class Schizomycetes, bacteria are now classified as prokaryotes. Unlike cells of animals and other eukaryotes, bacterial cells do not contain a nucleus and rarely harbour membrane-bound organelles. Although the term bacteria traditionally included all prokaryotes, the scientific classification changed after the discovery in the 1990s that prokaryotes consist of two very different groups of organisms that evolved from an ancient common ancestor. These evolutionary domains are called Bacteria and Archaea.[1]
Etymology
Orange labels: known ice ages.
Also see: Human timeline and Nature timeline
The word bacteria is the plural of the New Latin bacterium, which is the latinisation of the Greek βακτήριον (bakterion),[16] the diminutive of βακτηρία (bakteria), meaning "staff, cane",[17] because the first ones to be discovered were rod-shaped.[18][19]
Origin and early evolution
The ancestors of modern bacteria were unicellular microorganisms that were the first forms of life to appear on Earth, about 4 billion years ago. For about 3 billion years, most organisms were microscopic, and bacteria and archaea were the dominant forms of life.[20][21] In 2008, fossils of macroorganisms were discovered and named as the Francevillian biota. Although bacterial fossils exist, such as stromatolites, their lack of distinctive morphology prevents them from being used to examine the history of bacterial evolution, or to date the time of origin of a particular bacterial species. However, gene sequences can be used to reconstruct the bacterial phylogeny, and these studies indicate that bacteria diverged first from the archaeal/eukaryotic lineage.[22] Bacteria were also involved in the second great evolutionary divergence, that of the archaea and eukaryotes. Here, eukaryotes resulted from the entering of ancient bacteria into endosymbiotic associations with the ancestors of eukaryotic cells, which were themselves possibly related to the Archaea.[23][24] This involved the engulfment by proto-eukaryotic cells of alphaproteobacterial symbionts to form either mitochondria or hydrogenosomes, which are still found in all known Eukarya (sometimes in highly reduced form, e.g. in ancient "amitochondrial" protozoa). Later on, some eukaryotes that already contained mitochondria also engulfed cyanobacterial-like organisms. This led to the formation of chloroplasts in algae and plants. There are also some algae that originated from even later endosymbiotic events. Here, eukaryotes engulfed a eukaryotic algae that developed into a "second-generation" plastid.[25][26] This is known as secondary endosymbiosis.
In March 2017, researchers reported evidence of possibly the oldest forms of life on Earth. Putative fossilized microorganisms were discovered in hydrothermal vent precipitates in the Nuvvuagittuq Belt of Quebec, Canada, that may have lived as early as 4.280 billion years ago, not long after the oceans formed 4.4 billion years ago, and not long after the formation of the Earth 4.54 billion years ago.[27][28][29] According to biologist Stephen Blair Hedges, "If life arose relatively quickly on Earth … then it could be common in the universe."[30]
Morphology

Bacteria display a wide diversity of shapes and sizes, called morphologies. Bacterial cells are about one-tenth the size of eukaryotic cells and are typically 0.5–5.0 micrometres in length. However, a few species are visible to the unaided eye—for example, Thiomargarita namibiensis is up to half a millimetre long[31] and Epulopiscium fishelsoni reaches 0.7 mm.[32] Among the smallest bacteria are members of the genus Mycoplasma, which measure only 0.3 micrometres, as small as the largest viruses.[33] Some bacteria may be even smaller, but these ultramicrobacteria are not well-studied.[34]
Most bacterial species are either spherical, called cocci (sing. coccus, from Greek kókkos, grain, seed), or rod-shaped, called bacilli (sing. bacillus, from Latin baculus, stick). Elongation is associated with swimming.[35] Some bacteria, called vibrio, are shaped like slightly curved rods or comma-shaped; others can be spiral-shaped, called spirilla, or tightly coiled, called spirochaetes. A small number of species even have tetrahedral or cuboidal shapes.[36] More recently, some bacteria were discovered deep under Earth's crust that grow as branching filamentous types with a star-shaped cross-section. The large surface area to volume ratio of this morphology may give these bacteria an advantage in nutrient-poor environments.[37] This wide variety of shapes is determined by the bacterial cell wall and cytoskeleton, and is important because it can influence the ability of bacteria to acquire nutrients, attach to surfaces, swim through liquids and escape predators.[38][39]

Many bacterial species exist simply as single cells, others associate in characteristic patterns: Neisseria form diploids (pairs), Streptococcus form chains, and Staphylococcus group together in "bunch of grapes" clusters. Bacteria can also be elongated to form filaments, for example the Actinobacteria. Filamentous bacteria are often surrounded by a sheath that contains many individual cells. Certain types, such as species of the genus Nocardia, even form complex, branched filaments, similar in appearance to fungal mycelia.[40]
Bacteria often attach to surfaces and form dense aggregations in bacterial mats called biofilms. These biofilms can range from a few micrometres in thickness to up to half a metre in depth, and may contain multiple species of bacteria, protists and archaea. Bacteria living in biofilms display a complex arrangement of cells and extracellular components, forming secondary structures, such as microcolonies, through which there are networks of channels to enable better diffusion of nutrients.[41][42] In natural environments, such as soil or the surfaces of plants, the majority of bacteria are bound to surfaces in biofilms.[43] Biofilms are also important in medicine, as these structures are often present during chronic bacterial infections or in infections of implanted medical devices, and bacteria protected within biofilms are much harder to kill than individual isolated bacteria.[44]
Even more complex morphological changes are sometimes possible. For example, when starved of amino acids, Myxobacteria detect surrounding cells in a process known as quorum sensing, migrate towards each other, and aggregate to form fruiting bodies up to 500 micrometres long and containing approximately 100,000 bacterial cells.[45] In these fruiting bodies, the bacteria perform separate tasks; this type of cooperation is a simple type of multicellular organisation. For example, about one in 10 cells migrate to the top of these fruiting bodies and differentiate into a specialised dormant state called myxospores, which are more resistant to drying and other adverse environmental conditions than are ordinary cells.[46]
Cellular structure

Intracellular structures
The bacterial cell is surrounded by a cell membrane (also known as a lipid, cytoplasmic or plasma membrane). This membrane encloses the contents of the cell and acts as a barrier to hold nutrients, proteins and other essential components of the cytoplasm within the cell. As they are prokaryotes, bacteria do not usually have membrane-bound organelles in their cytoplasm, and thus contain few large intracellular structures. They lack a true nucleus, mitochondria, chloroplasts and the other organelles present in eukaryotic cells.[47] Bacteria were once seen as simple bags of cytoplasm, but structures such as the prokaryotic cytoskeleton[48][49] and the localisation of proteins to specific locations within the cytoplasm[48] that give bacteria some complexity have been discovered. These subcellular levels of organisation have been called "bacterial hyperstructures".[50]
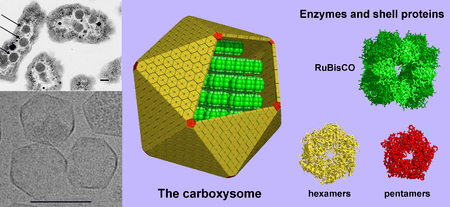
Bacterial microcompartments, such as carboxysomes,[51] provide a further level of organisation; they are compartments within bacteria that are surrounded by polyhedral protein shells, rather than by lipid membranes.[52] These "polyhedral organelles" localise and compartmentalise bacterial metabolism, a function performed by the membrane-bound organelles in eukaryotes.[53][54]
Many important biochemical reactions, such as energy generation, use concentration gradients across membranes. The general lack of internal membranes in bacteria means reactions such as electron transport occur across the cell membrane between the cytoplasm and the periplasmic space.[55] However, in many photosynthetic bacteria the plasma membrane is highly folded and fills most of the cell with layers of light-gathering membrane.[56] These light-gathering complexes may even form lipid-enclosed structures called chlorosomes in green sulfur bacteria.[57] Other proteins import nutrients across the cell membrane, or expel undesired molecules from the cytoplasm.
Bacteria do not have a membrane-bound nucleus, and their genetic material is typically a single circular bacterial chromosome of DNA located in the cytoplasm in an irregularly shaped body called the nucleoid.[59] The nucleoid contains the chromosome with its associated proteins and RNA. The phylum Planctomycetes[60] and candidate phylum Poribacteria[61] may be exceptions to the general absence of internal membranes in bacteria, because they appear to have a double membrane around their nucleoids and contain other membrane-bound cellular structures. Like all living organisms, bacteria contain ribosomes, often grouped in chains called polyribosomes, for the production of proteins, but the structure of the bacterial ribosome is different from that of eukaryotes and Archaea.[62] Bacterial ribosomes have a sedimentation rate of 70S (measured in Svedberg units): their subunits have rates of 30S and 50S. Some antibiotics bind specifically to 70S ribosomes and inhibit bacterial protein synthesis. Those antibiotics kill bacteria without affecting the larger 80S ribosomes of eukaryotic cells and without harming the host.
Some bacteria produce intracellular nutrient storage granules for later use, such as glycogen,[63] polyphosphate,[64] sulfur[65] or polyhydroxyalkanoates.[66] Certain bacterial species, such as the photosynthetic Cyanobacteria, produce internal gas vesicles, which they use to regulate their buoyancy—allowing them to move up or down into water layers with different light intensities and nutrient levels.[67] Intracellular membranes called chromatophores are also found in membranes of phototrophic bacteria. Used primarily for photosynthesis, they contain bacteriochlorophyll pigments and carotenoids. An early idea was that bacteria might contain membrane folds termed mesosomes, but these were later shown to be artefacts produced by the chemicals used to prepare the cells for electron microscopy. Inclusions are considered to be nonliving components of the cell that do not possess metabolic activity and are not bounded by membranes. The most common inclusions are glycogen, lipid droplets, crystals, and pigments. Volutin granules are cytoplasmic inclusions of complexed inorganic polyphosphate. These granules are called metachromatic granules due to their displaying the metachromatic effect; they appear red or blue when stained with the blue dyes methylene blue or toluidine blue. Gas vacuoles, which are freely permeable to gas, are membrane-bound vesicles present in some species of Cyanobacteria. They allow the bacteria to control their buoyancy. Microcompartments are widespread, membrane-bound organelles that are made of a protein shell that surrounds and encloses various enzymes. Carboxysomes are bacterial microcompartments that contain enzymes involved in carbon fixation. Magnetosomes are bacterial microcompartments, present in magnetotactic bacteria, that contain magnetic crystals.
Extracellular structures
In most bacteria, a cell wall is present on the outside of the cell membrane. The cell membrane and cell wall comprise the cell envelope. A common bacterial cell wall material is peptidoglycan (called "murein" in older sources), which is made from polysaccharide chains cross-linked by peptides containing D-amino acids.[68] Bacterial cell walls are different from the cell walls of plants and fungi, which are made of cellulose and chitin, respectively.[69] The cell wall of bacteria is also distinct from that of Archaea, which do not contain peptidoglycan. The cell wall is essential to the survival of many bacteria, and the antibiotic penicillin is able to kill bacteria by inhibiting a step in the synthesis of peptidoglycan.[69]
There are broadly speaking two different types of cell wall in bacteria, a thick one in the gram-positives and a thinner one in the gram-negatives. The names originate from the reaction of cells to the Gram stain, a long-standing test for the classification of bacterial species.[70]
Gram-positive bacteria possess a thick cell wall containing many layers of peptidoglycan and teichoic acids. In contrast, gram-negative bacteria have a relatively thin cell wall consisting of a few layers of peptidoglycan surrounded by a second lipid membrane containing lipopolysaccharides and lipoproteins. Lipopolysaccharides, also called endotoxins, are composed of polysaccharides and lipid A that is responsible for much of the toxicity of gram-negative bacteria. Most bacteria have the gram-negative cell wall, and only the Firmicutes and Actinobacteria have the alternative gram-positive arrangement.[71] These two groups were previously known as the low G+C and high G+C gram-positive bacteria, respectively. These differences in structure can produce differences in antibiotic susceptibility; for instance, vancomycin can kill only gram-positive bacteria and is ineffective against gram-negative pathogens, such as Haemophilus influenzae or Pseudomonas aeruginosa.[72] If the bacterial cell wall is entirely removed, it is called a protoplast, whereas if it is partially removed, it is called a spheroplast. β-Lactam antibiotics, such as penicillin, inhibit the formation of peptidoglycan cross-links in the bacterial cell wall. The enzyme lysozyme, found in human tears, also digests the cell wall of bacteria and is the body's main defence against eye infections.
Acid-fast bacteria, such as Mycobacteria, are resistant to decolorisation by acids during staining procedures. The high mycolic acid content of Mycobacteria is responsible for the staining pattern of poor absorption followed by high retention. The most common staining technique used to identify acid-fast bacteria is the Ziehl-Neelsen stain or acid-fast stain, in which the acid-fast bacilli are stained bright-red and stand out clearly against a blue background. L-form bacteria are strains of bacteria that lack cell walls. The main pathogenic bacteria in this class is Mycoplasma (not to be confused with Mycobacteria).
In many bacteria, an S-layer of rigidly arrayed protein molecules covers the outside of the cell.[73] This layer provides chemical and physical protection for the cell surface and can act as a macromolecular diffusion barrier. S-layers have diverse but mostly poorly understood functions, but are known to act as virulence factors in Campylobacter and contain surface enzymes in Bacillus stearothermophilus.[74]

Flagella are rigid protein structures, about 20 nanometres in diameter and up to 20 micrometres in length, that are used for motility. Flagella are driven by the energy released by the transfer of ions down an electrochemical gradient across the cell membrane.[75]
Fimbriae (sometimes called "attachment pili") are fine filaments of protein, usually 2–10 nanometres in diameter and up to several micrometres in length. They are distributed over the surface of the cell, and resemble fine hairs when seen under the electron microscope. Fimbriae are believed to be involved in attachment to solid surfaces or to other cells, and are essential for the virulence of some bacterial pathogens.[76] Pili (sing. pilus) are cellular appendages, slightly larger than fimbriae, that can transfer genetic material between bacterial cells in a process called conjugation where they are called conjugation pili or "sex pili" (see bacterial genetics, below).[77] They can also generate movement where they are called type IV pili (see movement, below).
Glycocalyx are produced by many bacteria to surround their cells, and vary in structural complexity: ranging from a disorganised slime layer of extra-cellular polymer to a highly structured capsule. These structures can protect cells from engulfment by eukaryotic cells such as macrophages (part of the human immune system).[78] They can also act as antigens and be involved in cell recognition, as well as aiding attachment to surfaces and the formation of biofilms.[79]
The assembly of these extracellular structures is dependent on bacterial secretion systems. These transfer proteins from the cytoplasm into the periplasm or into the environment around the cell. Many types of secretion systems are known and these structures are often essential for the virulence of pathogens, so are intensively studied.[80]
Endospores

Certain genera of gram-positive bacteria, such as Bacillus, Clostridium, Sporohalobacter, Anaerobacter, and Heliobacterium, can form highly resistant, dormant structures called endospores.[81] In almost all cases, one endospore is formed and this is not a reproductive process, although Anaerobacter can make up to seven endospores in a single cell.[82] Endospores have a central core of cytoplasm containing DNA and ribosomes surrounded by a cortex layer and protected by an impermeable and rigid coat. Dipicolinic acid is a chemical compound that composes 5% to 15% of the dry weight of bacterial spores. It is implicated as responsible for the heat resistance of the endospore.
Endospores show no detectable metabolism and can survive extreme physical and chemical stresses, such as high levels of UV light, gamma radiation, detergents, disinfectants, heat, freezing, pressure, and desiccation.[83] In this dormant state, these organisms may remain viable for millions of years,[84][85] and endospores even allow bacteria to survive exposure to the vacuum and radiation in space.[86] According to scientist Dr. Steinn Sigurdsson, "There are viable bacterial spores that have been found that are 40 million years old on Earth—and we know they're very hardened to radiation."[87] Endospore-forming bacteria can also cause disease: for example, anthrax can be contracted by the inhalation of Bacillus anthracis endospores, and contamination of deep puncture wounds with Clostridium tetani endospores causes tetanus.[88]
Metabolism
Bacteria exhibit an extremely wide variety of metabolic types.[89] The distribution of metabolic traits within a group of bacteria has traditionally been used to define their taxonomy, but these traits often do not correspond with modern genetic classifications.[90] Bacterial metabolism is classified into nutritional groups on the basis of three major criteria: the kind of energy used for growth, the source of carbon, and the electron donors used for growth. An additional criterion of respiratory microorganisms are the electron acceptors used for aerobic or anaerobic respiration.[91]
| Nutritional type | Source of energy | Source of carbon | Examples |
|---|---|---|---|
| Phototrophs | Sunlight | Organic compounds (photoheterotrophs) or carbon fixation (photoautotrophs) | Cyanobacteria, Green sulfur bacteria, Chloroflexi, or Purple bacteria |
| Lithotrophs | Inorganic compounds | Organic compounds (lithoheterotrophs) or carbon fixation (lithoautotrophs) | Thermodesulfobacteria, Hydrogenophilaceae, or Nitrospirae |
| Organotrophs | Organic compounds | Organic compounds (chemoheterotrophs) or carbon fixation (chemoautotrophs) | Bacillus, Clostridium or Enterobacteriaceae |
Carbon metabolism in bacteria is either heterotrophic, where organic carbon compounds are used as carbon sources, or autotrophic, meaning that cellular carbon is obtained by fixing carbon dioxide. Heterotrophic bacteria include parasitic types. Typical autotrophic bacteria are phototrophic cyanobacteria, green sulfur-bacteria and some purple bacteria, but also many chemolithotrophic species, such as nitrifying or sulfur-oxidising bacteria.[92] Energy metabolism of bacteria is either based on phototrophy, the use of light through photosynthesis, or based on chemotrophy, the use of chemical substances for energy, which are mostly oxidised at the expense of oxygen or alternative electron acceptors (aerobic/anaerobic respiration).

Bacteria are further divided into lithotrophs that use inorganic electron donors and organotrophs that use organic compounds as electron donors. Chemotrophic organisms use the respective electron donors for energy conservation (by aerobic/anaerobic respiration or fermentation) and biosynthetic reactions (e.g., carbon dioxide fixation), whereas phototrophic organisms use them only for biosynthetic purposes. Respiratory organisms use chemical compounds as a source of energy by taking electrons from the reduced substrate and transferring them to a terminal electron acceptor in a redox reaction. This reaction releases energy that can be used to synthesise ATP and drive metabolism. In aerobic organisms, oxygen is used as the electron acceptor. In anaerobic organisms other inorganic compounds, such as nitrate, sulfate or carbon dioxide are used as electron acceptors. This leads to the ecologically important processes of denitrification, sulfate reduction, and acetogenesis, respectively.
Another way of life of chemotrophs in the absence of possible electron acceptors is fermentation, wherein the electrons taken from the reduced substrates are transferred to oxidised intermediates to generate reduced fermentation products (e.g., lactate, ethanol, hydrogen, butyric acid). Fermentation is possible, because the energy content of the substrates is higher than that of the products, which allows the organisms to synthesise ATP and drive their metabolism.[93][94]
These processes are also important in biological responses to pollution; for example, sulfate-reducing bacteria are largely responsible for the production of the highly toxic forms of mercury (methyl- and dimethylmercury) in the environment.[95] Non-respiratory anaerobes use fermentation to generate energy and reducing power, secreting metabolic by-products (such as ethanol in brewing) as waste. Facultative anaerobes can switch between fermentation and different terminal electron acceptors depending on the environmental conditions in which they find themselves.
Lithotrophic bacteria can use inorganic compounds as a source of energy. Common inorganic electron donors are hydrogen, carbon monoxide, ammonia (leading to nitrification), ferrous iron and other reduced metal ions, and several reduced sulfur compounds. In unusual circumstances, the gas methane can be used by methanotrophic bacteria as both a source of electrons and a substrate for carbon anabolism.[96] In both aerobic phototrophy and chemolithotrophy, oxygen is used as a terminal electron acceptor, whereas under anaerobic conditions inorganic compounds are used instead. Most lithotrophic organisms are autotrophic, whereas organotrophic organisms are heterotrophic.
In addition to fixing carbon dioxide in photosynthesis, some bacteria also fix nitrogen gas using the enzyme nitrogenase. This environmentally important trait can be found in most bacteria of the metabolic types listed above.[97]
Regardless of the type of metabolic process they employ, the majority of bacteria are able to take in raw materials only in the form of relatively small molecules, which enter the cell by diffusion or through molecular channels in cell membranes. The Planctomycetes are the exception (as they are in possessing membranes around their nuclear material). It has recently been shown that Gemmata obscuriglobus is able to take in large molecules via a process that in some ways resembles endocytosis, the process used by eukaryotic cells to engulf external items.[32][98]
Growth and reproduction

Unlike in multicellular organisms, increases in cell size (cell growth) and reproduction by cell division are tightly linked in unicellular organisms. Bacteria grow to a fixed size and then reproduce through binary fission, a form of asexual reproduction.[99] Under optimal conditions, bacteria can grow and divide extremely rapidly, and bacterial populations can double as quickly as every 9.8 minutes.[100] In cell division, two identical clone daughter cells are produced. Some bacteria, while still reproducing asexually, form more complex reproductive structures that help disperse the newly formed daughter cells. Examples include fruiting body formation by Myxobacteria and aerial hyphae formation by Streptomyces, or budding. Budding involves a cell forming a protrusion that breaks away and produces a daughter cell.

In the laboratory, bacteria are usually grown using solid or liquid media. Solid growth media, such as agar plates, are used to isolate pure cultures of a bacterial strain. However, liquid growth media are used when measurement of growth or large volumes of cells are required. Growth in stirred liquid media occurs as an even cell suspension, making the cultures easy to divide and transfer, although isolating single bacteria from liquid media is difficult. The use of selective media (media with specific nutrients added or deficient, or with antibiotics added) can help identify specific organisms.[102]
Most laboratory techniques for growing bacteria use high levels of nutrients to produce large amounts of cells cheaply and quickly. However, in natural environments, nutrients are limited, meaning that bacteria cannot continue to reproduce indefinitely. This nutrient limitation has led the evolution of different growth strategies (see r/K selection theory). Some organisms can grow extremely rapidly when nutrients become available, such as the formation of algal (and cyanobacterial) blooms that often occur in lakes during the summer.[103] Other organisms have adaptations to harsh environments, such as the production of multiple antibiotics by Streptomyces that inhibit the growth of competing microorganisms.[104] In nature, many organisms live in communities (e.g., biofilms) that may allow for increased supply of nutrients and protection from environmental stresses.[43] These relationships can be essential for growth of a particular organism or group of organisms (syntrophy).[105]
Bacterial growth follows four phases. When a population of bacteria first enter a high-nutrient environment that allows growth, the cells need to adapt to their new environment. The first phase of growth is the lag phase, a period of slow growth when the cells are adapting to the high-nutrient environment and preparing for fast growth. The lag phase has high biosynthesis rates, as proteins necessary for rapid growth are produced.[106] The second phase of growth is the log phase, also known as the logarithmic or exponential phase. The log phase is marked by rapid exponential growth. The rate at which cells grow during this phase is known as the growth rate (k), and the time it takes the cells to double is known as the generation time (g). During log phase, nutrients are metabolised at maximum speed until one of the nutrients is depleted and starts limiting growth. The third phase of growth is the stationary phase and is caused by depleted nutrients. The cells reduce their metabolic activity and consume non-essential cellular proteins. The stationary phase is a transition from rapid growth to a stress response state and there is increased expression of genes involved in DNA repair, antioxidant metabolism and nutrient transport.[107] The final phase is the death phase where the bacteria run out of nutrients and die.
Genomes
The genomes of thousands of bacterial species have been sequenced, with at least 9,000 sequences completed and more than 42,000 left as "permanent" drafts (as of Sep 2016).[108]
Most bacteria have a single circular chromosome that can range in size from only 160,000 base pairs in the endosymbiotic bacteria Candidatus Carsonella ruddii,[109] to 12,200,000 base pairs in the soil-dwelling bacteria Sorangium cellulosum.[110] The genes in bacterial genomes are usually a single continuous stretch of DNA and although several different types of introns do exist in bacteria, these are much rarer than in eukaryotes.[111] Some bacteria, including the Spirochaetes of the genus Borrelia are a notable exception to this arrangement. Borrelia burgdorferi, the cause of Lyme disease, contains a single linear chromosome and several linear and circular plasmids.[112][113]
Plasmids are small extra-chromosomal DNAs that may contain genes for antibiotic resistance or virulence factors. Plasmids replicate independently of chromosomes, so it is possible that plasmids could be lost in bacterial cell division. Against this possibility is the fact that a single bacterium can contain hundreds of copies of a single plasmid.[114]
Genetics
Bacteria, as asexual organisms, inherit identical copies of their parent's genes (i.e., they are clonal). However, all bacteria can evolve by selection on changes to their genetic material DNA caused by genetic recombination or mutations. Mutations come from errors made during the replication of DNA or from exposure to mutagens. Mutation rates vary widely among different species of bacteria and even among different clones of a single species of bacteria.[115] Genetic changes in bacterial genomes come from either random mutation during replication or "stress-directed mutation", where genes involved in a particular growth-limiting process have an increased mutation rate.[116]
DNA transfer
Some bacteria also transfer genetic material between cells. This can occur in three main ways. First, bacteria can take up exogenous DNA from their environment, in a process called transformation. Genes can also be transferred by the process of transduction, when the integration of a bacteriophage introduces foreign DNA into the chromosome. The third method of gene transfer is conjugation, whereby DNA is transferred through direct cell contact.
Transduction of bacterial genes by bacteriophage appears to be a consequence of infrequent errors during intracellular assembly of virus particles, rather than a bacterial adaptation. Conjugation, in the much-studied E. coli system is determined by plasmid genes, and is an adaptation for transferring copies of the plasmid from one bacterial host to another. It is seldom that a conjugative plasmid integrates into the host bacterial chromosome, and subsequently transfers part of the host bacterial DNA to another bacterium. Plasmid-mediated transfer of host bacterial DNA also appears to be an accidental process rather than a bacterial adaptation.
Transformation, unlike transduction or conjugation, depends on numerous bacterial gene products that specifically interact to perform this complex process,[117] and thus transformation is clearly a bacterial adaptation for DNA transfer. In order for a bacterium to bind, take up and recombine donor DNA into its own chromosome, it must first enter a special physiological state termed competence (see Natural competence). In Bacillus subtilis, about 40 genes are required for the development of competence.[118] The length of DNA transferred during B. subtilis transformation can be between a third of a chromosome up to the whole chromosome.[119][120] Transformation appears to be common among bacterial species, and thus far at least 60 species are known to have the natural ability to become competent for transformation.[121] The development of competence in nature is usually associated with stressful environmental conditions, and seems to be an adaptation for facilitating repair of DNA damage in recipient cells.[122]
In ordinary circumstances, transduction, conjugation, and transformation involve transfer of DNA between individual bacteria of the same species, but occasionally transfer may occur between individuals of different bacterial species and this may have significant consequences, such as the transfer of antibiotic resistance.[123] In such cases, gene acquisition from other bacteria or the environment is called horizontal gene transfer and may be common under natural conditions.[124] Gene transfer is particularly important in antibiotic resistance as it allows the rapid transfer of resistance genes between different pathogens.[125]
Bacteriophages
Bacteriophages are viruses that infect bacteria. Many types of bacteriophage exist, some simply infect and lyse their host bacteria, while others insert into the bacterial chromosome. A bacteriophage can contain genes that contribute to its host's phenotype: for example, in the evolution of Escherichia coli O157:H7 and Clostridium botulinum, the toxin genes in an integrated phage converted a harmless ancestral bacterium into a lethal pathogen.[126] Bacteria resist phage infection through restriction modification systems that degrade foreign DNA,[127] and a system that uses CRISPR sequences to retain fragments of the genomes of phage that the bacteria have come into contact with in the past, which allows them to block virus replication through a form of RNA interference.[128][129] This CRISPR system provides bacteria with acquired immunity to infection.
Behaviour
Secretion
Bacteria frequently secrete chemicals into their environment in order to modify it favourably. The secretions are often proteins and may act as enzymes that digest some form of food in the environment.
Bioluminescence
A few bacteria have chemical systems that generate light. This bioluminescence often occurs in bacteria that live in association with fish, and the light probably serves to attract fish or other large animals.[130]
Multicellularity
Bacteria often function as multicellular aggregates known as biofilms, exchanging a variety of molecular signals for inter-cell communication, and engaging in coordinated multicellular behaviour.[131][132]
The communal benefits of multicellular cooperation include a cellular division of labour, accessing resources that cannot effectively be used by single cells, collectively defending against antagonists, and optimising population survival by differentiating into distinct cell types.[131] For example, bacteria in biofilms can have more than 500 times increased resistance to antibacterial agents than individual "planktonic" bacteria of the same species.[132]
One type of inter-cellular communication by a molecular signal is called quorum sensing, which serves the purpose of determining whether there is a local population density that is sufficiently high that it is productive to invest in processes that are only successful if large numbers of similar organisms behave similarly, as in excreting digestive enzymes or emitting light.
Quorum sensing allows bacteria to coordinate gene expression, and enables them to produce, release and detect autoinducers or pheromones which accumulate with the growth in cell population.[133]
Movement
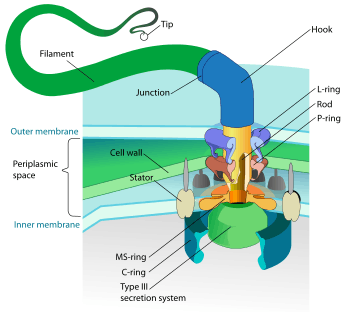
Many bacteria can move using a variety of mechanisms: flagella are used for swimming through fluids; bacterial gliding and twitching motility move bacteria across surfaces; and changes of buoyancy allow vertical motion.[134]
Swimming bacteria frequently move near 10 body lengths per second and a few as fast as 100. This makes them at least as fast as fish, on a relative scale.[135]
In bacterial gliding and twitching motility, bacteria use their type IV pili as a grappling hook, repeatedly extending it, anchoring it and then retracting it with remarkable force (>80 pN).[136]
Our observations redefine twitching motility as a rapid, highly organized mechanism of bacterial translocation by which Pseudomonas aeruginosa can disperse itself over large areas to colonize new territories. It is also now clear, both morphologically and genetically, that twitching motility and social gliding motility, such as occurs in Myxococcus xanthus, are essentially the same process.— Semmler, Whitchurch & Mattick (1999)[137]
Flagella are semi-rigid cylindrical structures that are rotated and function much like the propeller on a ship. Objects as small as bacteria operate a low Reynolds number and cylindrical forms are more efficient than the flat, paddle-like, forms appropriate at human-size scale.[138]
Bacterial species differ in the number and arrangement of flagella on their surface; some have a single flagellum (monotrichous), a flagellum at each end (amphitrichous), clusters of flagella at the poles of the cell (lophotrichous), while others have flagella distributed over the entire surface of the cell (peritrichous). The bacterial flagella is the best-understood motility structure in any organism and is made of about 20 proteins, with approximately another 30 proteins required for its regulation and assembly.[134] The flagellum is a rotating structure driven by a reversible motor at the base that uses the electrochemical gradient across the membrane for power.[139] This motor drives the motion of the filament, which acts as a propeller.
Many bacteria (such as E. coli) have two distinct modes of movement: forward movement (swimming) and tumbling. The tumbling allows them to reorient and makes their movement a three-dimensional random walk.[140] (See external links below for link to videos.) The flagella of a unique group of bacteria, the spirochaetes, are found between two membranes in the periplasmic space. They have a distinctive helical body that twists about as it moves.[134]
Motile bacteria are attracted or repelled by certain stimuli in behaviours called taxes: these include chemotaxis, phototaxis, energy taxis, and magnetotaxis.[141][142][143] In one peculiar group, the myxobacteria, individual bacteria move together to form waves of cells that then differentiate to form fruiting bodies containing spores.[46] The myxobacteria move only when on solid surfaces, unlike E. coli, which is motile in liquid or solid media.
Several Listeria and Shigella species move inside host cells by usurping the cytoskeleton, which is normally used to move organelles inside the cell. By promoting actin polymerisation at one pole of their cells, they can form a kind of tail that pushes them through the host cell's cytoplasm.[144]
Classification and identification

- ^ Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P (March 2006). "Toward automatic reconstruction of a highly resolved tree of life". Science. 311 (5765): 1283–7. Bibcode:2006Sci...311.1283C. PMID 16513982. doi:10.1126/science.1123061.
Classification seeks to describe the diversity of bacterial species by naming and grouping organisms based on similarities. Bacteria can be classified on the basis of cell structure, cellular metabolism or on differences in cell components, such as DNA, fatty acids, pigments, antigens and quinones.[102] While these schemes allowed the identification and classification of bacterial strains, it was unclear whether these differences represented variation between distinct species or between strains of the same species. This uncertainty was due to the lack of distinctive structures in most bacteria, as well as lateral gene transfer between unrelated species.[145] Due to lateral gene transfer, some closely related bacteria can have very different morphologies and metabolisms. To overcome this uncertainty, modern bacterial classification emphasises molecular systematics, using genetic techniques such as guanine cytosine ratio determination, genome-genome hybridisation, as well as sequencing genes that have not undergone extensive lateral gene transfer, such as the rRNA gene.[146] Classification of bacteria is determined by publication in the International Journal of Systematic Bacteriology,[147] and Bergey's Manual of Systematic Bacteriology.[148] The International Committee on Systematic Bacteriology (ICSB) maintains international rules for the naming of bacteria and taxonomic categories and for the ranking of them in the International Code of Nomenclature of Bacteria.
The term "bacteria" was traditionally applied to all microscopic, single-cell prokaryotes. However, molecular systematics showed prokaryotic life to consist of two separate domains, originally called Eubacteria and Archaebacteria, but now called Bacteria and Archaea that evolved independently from an ancient common ancestor.[1] The archaea and eukaryotes are more closely related to each other than either is to the bacteria. These two domains, along with Eukarya, are the basis of the three-domain system, which is currently the most widely used classification system in microbiology.[149] However, due to the relatively recent introduction of molecular systematics and a rapid increase in the number of genome sequences that are available, bacterial classification remains a changing and expanding field.[4][150] For example, a few biologists argue that the Archaea and Eukaryotes evolved from gram-positive bacteria.[151]
The identification of bacteria in the laboratory is particularly relevant in medicine, where the correct treatment is determined by the bacterial species causing an infection. Consequently, the need to identify human pathogens was a major impetus for the development of techniques to identify bacteria.
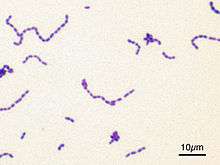
The Gram stain, developed in 1884 by Hans Christian Gram, characterises bacteria based on the structural characteristics of their cell walls.[70] The thick layers of peptidoglycan in the "gram-positive" cell wall stain purple, while the thin "gram-negative" cell wall appears pink. By combining morphology and Gram-staining, most bacteria can be classified as belonging to one of four groups (gram-positive cocci, gram-positive bacilli, gram-negative cocci and gram-negative bacilli). Some organisms are best identified by stains other than the Gram stain, particularly mycobacteria or Nocardia, which show acid-fastness on Ziehl–Neelsen or similar stains.[152] Other organisms may need to be identified by their growth in special media, or by other techniques, such as serology.
Culture techniques are designed to promote the growth and identify particular bacteria, while restricting the growth of the other bacteria in the sample. Often these techniques are designed for specific specimens; for example, a sputum sample will be treated to identify organisms that cause pneumonia, while stool specimens are cultured on selective media to identify organisms that cause diarrhoea, while preventing growth of non-pathogenic bacteria. Specimens that are normally sterile, such as blood, urine or spinal fluid, are cultured under conditions designed to grow all possible organisms.[102][153] Once a pathogenic organism has been isolated, it can be further characterised by its morphology, growth patterns (such as aerobic or anaerobic growth), patterns of hemolysis, and staining.
As with bacterial classification, identification of bacteria is increasingly using molecular methods. Diagnostics using DNA-based tools, such as polymerase chain reaction, are increasingly popular due to their specificity and speed, compared to culture-based methods.[154] These methods also allow the detection and identification of "viable but nonculturable" cells that are metabolically active but non-dividing.[155] However, even using these improved methods, the total number of bacterial species is not known and cannot even be estimated with any certainty. Following present classification, there are a little less than 9,300 known species of prokaryotes, which includes bacteria and archaea;[156] but attempts to estimate the true number of bacterial diversity have ranged from 107 to 109 total species—and even these diverse estimates may be off by many orders of magnitude.[157][158]
Interactions with other organisms
Despite their apparent simplicity, bacteria can form complex associations with other organisms. These symbiotic associations can be divided into parasitism, mutualism and commensalism. Due to their small size, commensal bacteria are ubiquitous and grow on animals and plants exactly as they will grow on any other surface. However, their growth can be increased by warmth and sweat, and large populations of these organisms in humans are the cause of body odour.
Predators
Some species of bacteria kill and then consume other microorganisms, these species are called predatory bacteria.[161] These include organisms such as Myxococcus xanthus, which forms swarms of cells that kill and digest any bacteria they encounter.[162] Other bacterial predators either attach to their prey in order to digest them and absorb nutrients, such as Vampirovibrio chlorellavorus,[163] or invade another cell and multiply inside the cytosol, such as Daptobacter.[164] These predatory bacteria are thought to have evolved from saprophages that consumed dead microorganisms, through adaptations that allowed them to entrap and kill other organisms.[165]
Mutualists
Certain bacteria form close spatial associations that are essential for their survival. One such mutualistic association, called interspecies hydrogen transfer, occurs between clusters of anaerobic bacteria that consume organic acids, such as butyric acid or propionic acid, and produce hydrogen, and methanogenic Archaea that consume hydrogen.[166] The bacteria in this association are unable to consume the organic acids as this reaction produces hydrogen that accumulates in their surroundings. Only the intimate association with the hydrogen-consuming Archaea keeps the hydrogen concentration low enough to allow the bacteria to grow.
In soil, microorganisms that reside in the rhizosphere (a zone that includes the root surface and the soil that adheres to the root after gentle shaking) carry out nitrogen fixation, converting nitrogen gas to nitrogenous compounds.[167] This serves to provide an easily absorbable form of nitrogen for many plants, which cannot fix nitrogen themselves. Many other bacteria are found as symbionts in humans and other organisms. For example, the presence of over 1,000 bacterial species in the normal human gut flora of the intestines can contribute to gut immunity, synthesise vitamins, such as folic acid, vitamin K and biotin, convert sugars to lactic acid (see Lactobacillus), as well as fermenting complex undigestible carbohydrates.[168][169][170] The presence of this gut flora also inhibits the growth of potentially pathogenic bacteria (usually through competitive exclusion) and these beneficial bacteria are consequently sold as probiotic dietary supplements.[171]
Pathogens

If bacteria form a parasitic association with other organisms, they are classed as pathogens. Pathogenic bacteria are a major cause of human death and disease and cause infections such as tetanus, typhoid fever, diphtheria, syphilis, cholera, foodborne illness, leprosy and tuberculosis. A pathogenic cause for a known medical disease may only be discovered many years after, as was the case with Helicobacter pylori and peptic ulcer disease. Bacterial diseases are also important in agriculture, with bacteria causing leaf spot, fire blight and wilts in plants, as well as Johne's disease, mastitis, salmonella and anthrax in farm animals.
Each species of pathogen has a characteristic spectrum of interactions with its human hosts. Some organisms, such as Staphylococcus or Streptococcus, can cause skin infections, pneumonia, meningitis and even overwhelming sepsis, a systemic inflammatory response producing shock, massive vasodilation and death.[172] Yet these organisms are also part of the normal human flora and usually exist on the skin or in the nose without causing any disease at all. Other organisms invariably cause disease in humans, such as the Rickettsia, which are obligate intracellular parasites able to grow and reproduce only within the cells of other organisms. One species of Rickettsia causes typhus, while another causes Rocky Mountain spotted fever. Chlamydia, another phylum of obligate intracellular parasites, contains species that can cause pneumonia, or urinary tract infection and may be involved in coronary heart disease.[173] Finally, some species, such as Pseudomonas aeruginosa, Burkholderia cenocepacia, and Mycobacterium avium, are opportunistic pathogens and cause disease mainly in people suffering from immunosuppression or cystic fibrosis.[174][175]
Bacterial infections may be treated with antibiotics, which are classified as bacteriocidal if they kill bacteria, or bacteriostatic if they just prevent bacterial growth. There are many types of antibiotics and each class inhibits a process that is different in the pathogen from that found in the host. An example of how antibiotics produce selective toxicity are chloramphenicol and puromycin, which inhibit the bacterial ribosome, but not the structurally different eukaryotic ribosome.[176] Antibiotics are used both in treating human disease and in intensive farming to promote animal growth, where they may be contributing to the rapid development of antibiotic resistance in bacterial populations.[177] Infections can be prevented by antiseptic measures such as sterilising the skin prior to piercing it with the needle of a syringe, and by proper care of indwelling catheters. Surgical and dental instruments are also sterilised to prevent contamination by bacteria. Disinfectants such as bleach are used to kill bacteria or other pathogens on surfaces to prevent contamination and further reduce the risk of infection.
Significance in technology and industry
Bacteria, often lactic acid bacteria, such as Lactobacillus and Lactococcus, in combination with yeasts and moulds, have been used for thousands of years in the preparation of fermented foods, such as cheese, pickles, soy sauce, sauerkraut, vinegar, wine and yogurt.[178][179]
The ability of bacteria to degrade a variety of organic compounds is remarkable and has been used in waste processing and bioremediation. Bacteria capable of digesting the hydrocarbons in petroleum are often used to clean up oil spills.[180] Fertiliser was added to some of the beaches in Prince William Sound in an attempt to promote the growth of these naturally occurring bacteria after the 1989 Exxon Valdez oil spill. These efforts were effective on beaches that were not too thickly covered in oil. Bacteria are also used for the bioremediation of industrial toxic wastes.[181] In the chemical industry, bacteria are most important in the production of enantiomerically pure chemicals for use as pharmaceuticals or agrichemicals.[182]
Bacteria can also be used in the place of pesticides in the biological pest control. This commonly involves Bacillus thuringiensis (also called BT), a gram-positive, soil dwelling bacterium. Subspecies of this bacteria are used as a Lepidopteran-specific insecticides under trade names such as Dipel and Thuricide.[183] Because of their specificity, these pesticides are regarded as environmentally friendly, with little or no effect on humans, wildlife, pollinators and most other beneficial insects.[184][185]
Because of their ability to quickly grow and the relative ease with which they can be manipulated, bacteria are the workhorses for the fields of molecular biology, genetics and biochemistry. By making mutations in bacterial DNA and examining the resulting phenotypes, scientists can determine the function of genes, enzymes and metabolic pathways in bacteria, then apply this knowledge to more complex organisms.[186] This aim of understanding the biochemistry of a cell reaches its most complex expression in the synthesis of huge amounts of enzyme kinetic and gene expression data into mathematical models of entire organisms. This is achievable in some well-studied bacteria, with models of Escherichia coli metabolism now being produced and tested.[187][188] This understanding of bacterial metabolism and genetics allows the use of biotechnology to bioengineer bacteria for the production of therapeutic proteins, such as insulin, growth factors, or antibodies.[189][190]
Because of their importance for research in general, samples of bacterial strains are isolated and preserved in Biological Resource Centers. This ensures the availability of the strain to scientists worldwide.
History of bacteriology
._Natuurkundige_te_Delft_Rijksmuseum_SK-A-957.jpeg)
Bacteria were first observed by the Dutch microscopist Antonie van Leeuwenhoek in 1676, using a single-lens microscope of his own design.[191] He then published his observations in a series of letters to the Royal Society of London.[192][193][194] Bacteria were Leeuwenhoek's most remarkable microscopic discovery. They were just at the limit of what his simple lenses could make out and, in one of the most striking hiatuses in the history of science, no one else would see them again for over a century.[195] Only then were his by-then-largely-forgotten observations of bacteria—as opposed to his famous "animalcules" (spermatozoa)—taken seriously.
Christian Gottfried Ehrenberg introduced the word "bacterium" in 1828.[196] In fact, his Bacterium was a genus that contained non-spore-forming rod-shaped bacteria,[197] as opposed to Bacillus, a genus of spore-forming rod-shaped bacteria defined by Ehrenberg in 1835.[198]
Louis Pasteur demonstrated in 1859 that the growth of microorganisms causes the fermentation process, and that this growth is not due to spontaneous generation. (Yeasts and moulds, commonly associated with fermentation, are not bacteria, but rather fungi.) Along with his contemporary Robert Koch, Pasteur was an early advocate of the germ theory of disease.[199]
Robert Koch, a pioneer in medical microbiology, worked on cholera, anthrax and tuberculosis. In his research into tuberculosis Koch finally proved the germ theory, for which he received a Nobel Prize in 1905.[200] In Koch's postulates, he set out criteria to test if an organism is the cause of a disease, and these postulates are still used today.[201]
Though it was known in the nineteenth century that bacteria are the cause of many diseases, no effective antibacterial treatments were available.[202] In 1910, Paul Ehrlich developed the first antibiotic, by changing dyes that selectively stained Treponema pallidum—the spirochaete that causes syphilis—into compounds that selectively killed the pathogen.[203] Ehrlich had been awarded a 1908 Nobel Prize for his work on immunology, and pioneered the use of stains to detect and identify bacteria, with his work being the basis of the Gram stain and the Ziehl–Neelsen stain.[204]
A major step forward in the study of bacteria came in 1977 when Carl Woese recognised that archaea have a separate line of evolutionary descent from bacteria.[2] This new phylogenetic taxonomy depended on the sequencing of 16S ribosomal RNA, and divided prokaryotes into two evolutionary domains, as part of the three-domain system.[1]
See also
- Bacteriotherapy
- Genetically modified bacteria
- List of bacterial orders
- Panspermia
- Polysaccharide encapsulated bacteria
- Psychrotrophic bacteria
References
- 1 2 3 4 Woese CR, Kandler O, Wheelis ML (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proceedings of the National Academy of Sciences of the United States of America. 87 (12): 4576–9. Bibcode:1990PNAS...87.4576W. PMC 54159
 . PMID 2112744. doi:10.1073/pnas.87.12.4576.
. PMID 2112744. doi:10.1073/pnas.87.12.4576. - 1 2 Woese CR, Fox GE (1977). "Phylogenetic structure of the prokaryotic domain: the primary kingdoms". Proceedings of the National Academy of Sciences of the United States of America. 74 (11): 5088–90. Bibcode:1977PNAS...74.5088W. PMC 432104
 . PMID 270744. doi:10.1073/pnas.74.11.5088.
. PMID 270744. doi:10.1073/pnas.74.11.5088. - ↑ Fredrickson JK, Zachara JM, Balkwill DL, Kennedy D, Li SM, Kostandarithes HM, Daly MJ, Romine MF, Brockman FJ (2004). "Geomicrobiology of high-level nuclear waste-contaminated vadose sediments at the Hanford site, Washington state". Applied and Environmental Microbiology. 70 (7): 4230–41. PMC 444790
 . PMID 15240306. doi:10.1128/AEM.70.7.4230-4241.2004.
. PMID 15240306. doi:10.1128/AEM.70.7.4230-4241.2004. - 1 2 Rappé MS, Giovannoni SJ (2003). "The uncultured microbial majority". Annual Review of Microbiology. 57: 369–94. PMID 14527284. doi:10.1146/annurev.micro.57.030502.090759.
- ↑ Whitman WB, Coleman DC, Wiebe WJ (1998). "Prokaryotes: the unseen majority". Proceedings of the National Academy of Sciences of the United States of America. 95 (12): 6578–83. Bibcode:1998PNAS...95.6578W. PMC 33863
 . PMID 9618454. doi:10.1073/pnas.95.12.6578.
. PMID 9618454. doi:10.1073/pnas.95.12.6578. - ↑ C.Michael Hogan. 2010. Bacteria. Encyclopedia of Earth. eds. Sidney Draggan and C.J.Cleveland, National Council for Science and the Environment, Washington DC Archived 11 May 2011 at the Wayback Machine.
- ↑ Forbes, S.L. (2008). "Decomposition Chemistry in a Burial Environment". In M. Tibbett; D.O. Carter. Soil Analysis in Forensic Taphonomy. CRC Press. pp. 203–223. ISBN 1-4200-6991-8.
- 1 2 3 Choi CQ (17 March 2013). "Microbes Thrive in Deepest Spot on Earth". LiveScience. Retrieved 17 March 2013.
- ↑ Glud R, Wenzhöfer F, Middelboe M, Oguri K, Turnewitsch R, Canfield DE, Kitazato H (2013). "High rates of microbial carbon turnover in sediments in the deepest oceanic trench on Earth". Nature Geoscience. 6 (4): 284–288. Bibcode:2013NatGe...6..284G. doi:10.1038/ngeo1773.
- ↑ Oskin B (14 March 2013). "Intraterrestrials: Life Thrives in Ocean Floor". LiveScience. Retrieved 17 March 2013.
- ↑ Sender R, Fuchs S, Milo R (2016). "Revised estimates for the number of human and bacteria cells in the body". bioRxiv 036103
 .
. - ↑ Sears CL (2005). "A dynamic partnership: celebrating our gut flora". Anaerobe. 11 (5): 247–51. PMID 16701579. doi:10.1016/j.anaerobe.2005.05.001.
- ↑ "2002 WHO mortality data". Retrieved 20 January 2007.
- ↑ "Metal-Mining Bacteria Are Green Chemists". Science Daily. 2 September 2010.
- ↑ Ishige T, Honda K, Shimizu S (2005). "Whole organism biocatalysis". Current Opinion in Chemical Biology. 9 (2): 174–80. PMID 15811802. doi:10.1016/j.cbpa.2005.02.001.
- ↑ βακτήριον. Liddell, Henry George; Scott, Robert; A Greek–English Lexicon at the Perseus Project.
- ↑ βακτηρία in Liddell and Scott.
- ↑ bacterium, on Oxford Dictionaries.
- ↑ Harper, Douglas. "bacteria". Online Etymology Dictionary.
- ↑ Schopf JW (1994). "Disparate rates, differing fates: tempo and mode of evolution changed from the Precambrian to the Phanerozoic". Proceedings of the National Academy of Sciences of the United States of America. 91 (15): 6735–42. Bibcode:1994PNAS...91.6735S. PMC 44277
 . PMID 8041691. doi:10.1073/pnas.91.15.6735.
. PMID 8041691. doi:10.1073/pnas.91.15.6735. - ↑ DeLong EF, Pace NR (2001). "Environmental diversity of bacteria and archaea". Syst Biol. 50 (4): 470–8. PMID 12116647. doi:10.1080/106351501750435040.
- ↑ Brown JR, Doolittle WF (1997). "Archaea and the prokaryote-to-eukaryote transition". Microbiology and Molecular Biology Reviews. 61 (4): 456–502. PMC 232621
 . PMID 9409149.
. PMID 9409149. - ↑ Poole AM, Penny D (2007). "Evaluating hypotheses for the origin of eukaryotes". BioEssays. 29 (1): 74–84. PMID 17187354. doi:10.1002/bies.20516.
- ↑ Dyall SD, Brown MT, Johnson PJ (2004). "Ancient invasions: from endosymbionts to organelles". Science. 304 (5668): 253–7. Bibcode:2004Sci...304..253D. PMID 15073369. doi:10.1126/science.1094884.
- ↑ Lang BF, Gray MW, Burger G (1999). "Mitochondrial genome evolution and the origin of eukaryotes". Annu Rev Genet. 33: 351–97. PMID 10690412. doi:10.1146/annurev.genet.33.1.351.
- ↑ McFadden GI (1999). "Endosymbiosis and evolution of the plant cell". Current Opinion in Plant Biology. 2 (6): 513–9. PMID 10607659. doi:10.1016/S1369-5266(99)00025-4.
- ↑ Dodd, Matthew S.; Papineau, Dominic; Grenne, Tor; Slack, John F.; Rittner, Martin; Pirajno, Franco; O'Neil, Jonathan; Little, Crispin T. S. (1 March 2017). "Evidence for early life in Earth’s oldest hydrothermal vent precipitates". Nature. 543: 60–64. doi:10.1038/nature21377. Retrieved 2 March 2017.
- ↑ Zimmer, Carl (1 March 2017). "Scientists Say Canadian Bacteria Fossils May Be Earth’s Oldest". New York Times. Retrieved 2 March 2017.
- ↑ Ghosh, Pallab (1 March 2017). "Earliest evidence of life on Earth 'found'". BBC News. Retrieved 2 March 2017.
- ↑ Borenstein, Seth (19 October 2015). "Hints of life on what was thought to be desolate early Earth". Excite. Yonkers, NY: Mindspark Interactive Network. Associated Press. Retrieved 2015-10-20.
- ↑ Schulz HN, Jorgensen BB (2001). "Big bacteria". Annu Rev Microbiol. 55: 105–37. PMID 11544351. doi:10.1146/annurev.micro.55.1.105.
- 1 2 Williams C (2011). "Who are you calling simple?". New Scientist. 211 (2821): 38–41. doi:10.1016/S0262-4079(11)61709-0.
- ↑ Robertson J, Gomersall M, Gill P (1975). "Mycoplasma hominis: growth, reproduction, and isolation of small viable cells". J Bacteriol. 124 (2): 1007–18. PMC 235991
 . PMID 1102522.
. PMID 1102522. - ↑ Velimirov B (2001). "Nanobacteria, Ultramicrobacteria and Starvation Forms: A Search for the Smallest Metabolizing Bacterium". Microbes and Environments. 16 (2): 67–77. doi:10.1264/jsme2.2001.67.
- ↑ Dusenbery, David B. (2009). Living at Micro Scale, pp. 20–25. Harvard University Press, Cambridge, Mass. ISBN 978-0-674-03116-6.
- ↑ Fritz I, Strömpl C, Abraham WR (2004). "Phylogenetic relationships of the genera Stella, Labrys and Angulomicrobium within the 'Alphaproteobacteria' and description of Angulomicrobium amanitiforme sp. nov". Int J Syst Evol Microbiol. 54 (Pt 3): 651–7. PMID 15143003. doi:10.1099/ijs.0.02746-0.
- ↑ Wanger G, Onstott TC, Southam G (2008). "Stars of the terrestrial deep subsurface: A novel 'star-shaped' bacterial morphotype from a South African platinum mine". Geobiology. 6 (3): 325–30. PMID 18498531. doi:10.1111/j.1472-4669.2008.00163.x.
- ↑ Cabeen MT, Jacobs-Wagner C (2005). "Bacterial cell shape". Nature Reviews Microbiology. 3 (8): 601–10. PMID 16012516. doi:10.1038/nrmicro1205.
- ↑ Young KD (2006). "The selective value of bacterial shape". Microbiol Mol Biol Rev. 70 (3): 660–703. PMC 1594593
 . PMID 16959965. doi:10.1128/MMBR.00001-06.
. PMID 16959965. doi:10.1128/MMBR.00001-06. - ↑ Douwes KE, Schmalzbauer E, Linde HJ, Reisberger EM, Fleischer K, Lehn N, Landthaler M, Vogt T (2003). "Branched filaments no fungus, ovoid bodies no bacteria: Two unusual cases of mycetoma". J Am Acad Dermatol. 49 (2 Suppl Case Reports): S170–3. PMID 12894113. doi:10.1067/mjd.2003.302.
- ↑ Donlan RM (2002). "Biofilms: microbial life on surfaces". Emerg Infect Dis. 8 (9): 881–90. PMC 2732559
 . PMID 12194761. doi:10.3201/eid0809.020063.
. PMID 12194761. doi:10.3201/eid0809.020063. - ↑ Branda SS, Vik S, Friedman L, Kolter R (2005). "Biofilms: the matrix revisited". Trends Microbiol. 13 (1): 20–6. PMID 15639628. doi:10.1016/j.tim.2004.11.006.
- 1 2 Davey ME, O'toole GA (2000). "Microbial biofilms: from ecology to molecular genetics". Microbiol Mol Biol Rev. 64 (4): 847–67. PMC 99016
 . PMID 11104821. doi:10.1128/MMBR.64.4.847-867.2000.
. PMID 11104821. doi:10.1128/MMBR.64.4.847-867.2000. - ↑ Donlan RM, Costerton JW (2002). "Biofilms: survival mechanisms of clinically relevant microorganisms". Clin Microbiol Rev. 15 (2): 167–93. PMC 118068
 . PMID 11932229. doi:10.1128/CMR.15.2.167-193.2002.
. PMID 11932229. doi:10.1128/CMR.15.2.167-193.2002. - ↑ Shimkets LJ (1999). "Intercellular signaling during fruiting-body development of Myxococcus xanthus". Annu Rev Microbiol. 53: 525–49. PMID 10547700. doi:10.1146/annurev.micro.53.1.525.
- 1 2 Kaiser D (2004). "Signaling in myxobacteria". Annu Rev Microbiol. 58: 75–98. PMID 15487930. doi:10.1146/annurev.micro.58.030603.123620.
- ↑ Berg JM, Tymoczko JL, Stryer L (2002). Molecular Cell Biology (5th ed.). WH Freeman. ISBN 0-7167-4955-6.
- 1 2 Gitai Z (2005). "The new bacterial cell biology: moving parts and subcellular architecture". Cell. 120 (5): 577–86. PMID 15766522. doi:10.1016/j.cell.2005.02.026.
- ↑ Shih YL, Rothfield L (2006). "The bacterial cytoskeleton". Microbiology and Molecular Biology Reviews. 70 (3): 729–54. PMC 1594594
 . PMID 16959967. doi:10.1128/MMBR.00017-06.
. PMID 16959967. doi:10.1128/MMBR.00017-06. - ↑ Norris V, den Blaauwen T, Cabin-Flaman A, Doi RH, Harshey R, Janniere L, Jimenez-Sanchez A, Jin DJ, Levin PA, Mileykovskaya E, Minsky A, Saier M, Skarstad K (2007). "Functional taxonomy of bacterial hyperstructures". Microbiology and Molecular Biology Reviews. 71 (1): 230–53. PMC 1847379
 . PMID 17347523. doi:10.1128/MMBR.00035-06.
. PMID 17347523. doi:10.1128/MMBR.00035-06. - ↑ Kerfeld CA, Sawaya MR, Tanaka S, Nguyen CV, Phillips M, Beeby M, Yeates TO (2005). "Protein structures forming the shell of primitive bacterial organelles". Science. 309 (5736): 936–8. Bibcode:2005Sci...309..936K. PMID 16081736. doi:10.1126/science.1113397.
- ↑ Bobik TA (2007). "Bacterial Microcompartments" (PDF). Microbe. Am Soc Microbiol. 2: 25–31. doi:10.1128/microbe.2.25.1. Archived from the original (PDF) on 2 August 2008.
- ↑ Bobik TA (2006). "Polyhedral organelles compartmenting bacterial metabolic processes". Applied Microbiology and Biotechnology. 70 (5): 517–25. PMID 16525780. doi:10.1007/s00253-005-0295-0.
- ↑ Yeates TO, Kerfeld CA, Heinhorst S, Cannon GC, Shively JM (2008). "Protein-based organelles in bacteria: carboxysomes and related microcompartments". Nature Reviews Microbiology. 6 (9): 681–91. PMID 18679172. doi:10.1038/nrmicro1913.
- ↑ Harold FM (1972). "Conservation and transformation of energy by bacterial membranes". Bacteriological Reviews. 36 (2): 172–230. PMC 408323
 . PMID 4261111.
. PMID 4261111. - ↑ Bryant DA, Frigaard NU (2006). "Prokaryotic photosynthesis and phototrophy illuminated". Trends Microbiol. 14 (11): 488–96. PMID 16997562. doi:10.1016/j.tim.2006.09.001.
- ↑ Psencík J, Ikonen TP, Laurinmäki P, Merckel MC, Butcher SJ, Serimaa RE, Tuma R (2004). "Lamellar organization of pigments in chlorosomes, the light harvesting complexes of green photosynthetic bacteria". Biophys. J. 87 (2): 1165–72. Bibcode:2004BpJ....87.1165P. PMC 1304455
 . PMID 15298919. doi:10.1529/biophysj.104.040956.
. PMID 15298919. doi:10.1529/biophysj.104.040956. - ↑ Tanaka S, Kerfeld CA, Sawaya MR, Cai F, Heinhorst S, Cannon GC, Yeates TO (2008). "Atomic-level models of the bacterial carboxysome shell". Science. 319 (5866): 1083–6. Bibcode:2008Sci...319.1083T. PMID 18292340. doi:10.1126/science.1151458.
- ↑ Thanbichler M, Wang SC, Shapiro L (2005). "The bacterial nucleoid: a highly organized and dynamic structure". J Cell Biochem. 96 (3): 506–21. PMID 15988757. doi:10.1002/jcb.20519.
- ↑ Fuerst JA (2005). "Intracellular compartmentation in planctomycetes". Annu Rev Microbiol. 59: 299–328. PMID 15910279. doi:10.1146/annurev.micro.59.030804.121258.
- ↑ Fieseler L, Horn M, Wagner M, Hentschel U (June 2004). "Discovery of the novel candidate phylum "Poribacteria" in marine sponges". Applied and Environmental Microbiology. 70 (6): 3724–32. PMC 427773
 . PMID 15184179. doi:10.1128/AEM.70.6.3724-3732.2004.
. PMID 15184179. doi:10.1128/AEM.70.6.3724-3732.2004. - ↑ Poehlsgaard J, Douthwaite S (2005). "The bacterial ribosome as a target for antibiotics". Nature Reviews Microbiology. 3 (11): 870–81. PMID 16261170. doi:10.1038/nrmicro1265.
- ↑ Yeo M, Chater K (2005). "The interplay of glycogen metabolism and differentiation provides an insight into the developmental biology of Streptomyces coelicolor". Microbiology. 151 (Pt 3): 855–61. PMID 15758231. doi:10.1099/mic.0.27428-0.
- ↑ Shiba T, Tsutsumi K, Ishige K, Noguchi T (2000). "Inorganic polyphosphate and polyphosphate kinase: their novel biological functions and applications". Biochemistry (Mosc). 65 (3): 315–23. PMID 10739474.
- ↑ Brune DC (1995). "Isolation and characterization of sulfur globule proteins from Chromatium vinosum and Thiocapsa roseopersicina". Archives of Microbiology. 163 (6): 391–9. PMID 7575095. doi:10.1007/BF00272127.
- ↑ Kadouri D, Jurkevitch E, Okon Y, Castro-Sowinski S (2005). "Ecological and agricultural significance of bacterial polyhydroxyalkanoates". Critical Reviews in Microbiology. 31 (2): 55–67. PMID 15986831. doi:10.1080/10408410590899228.
- ↑ Walsby AE (1994). "Gas vesicles". Microbiological Reviews. 58 (1): 94–144. PMC 372955
 . PMID 8177173.
. PMID 8177173. - ↑ van Heijenoort J (2001). "Formation of the glycan chains in the synthesis of bacterial peptidoglycan". Glycobiology. 11 (3): 25R–36R. PMID 11320055. doi:10.1093/glycob/11.3.25R.
- 1 2 Koch AL (2003). "Bacterial wall as target for attack: past, present, and future research". Clin Microbiol Rev. 16 (4): 673–87. PMC 207114
 . PMID 14557293. doi:10.1128/CMR.16.4.673-687.2003.
. PMID 14557293. doi:10.1128/CMR.16.4.673-687.2003. - 1 2 Gram, HC (1884). "Über die isolierte Färbung der Schizomyceten in Schnitt- und Trockenpräparaten". Fortschr. Med. 2: 185–189.
- ↑ Hugenholtz P (2002). "Exploring prokaryotic diversity in the genomic era". Genome Biology. 3 (2): reviews0003.1–reviews0003.8. PMC 139013
 . PMID 11864374. doi:10.1186/gb-2002-3-2-reviews0003.
. PMID 11864374. doi:10.1186/gb-2002-3-2-reviews0003. - ↑ Walsh FM, Amyes SG (2004). "Microbiology and drug resistance mechanisms of fully resistant pathogens". Current Opinion in Microbiology. 7 (5): 439–44. PMID 15451497. doi:10.1016/j.mib.2004.08.007.
- ↑ Engelhardt H, Peters J (1998). "Structural research on surface layers: a focus on stability, surface layer homology domains, and surface layer-cell wall interactions". J Struct Biol. 124 (2–3): 276–302. PMID 10049812. doi:10.1006/jsbi.1998.4070.
- ↑ Beveridge TJ, Pouwels PH, Sára M, Kotiranta A, Lounatmaa K, Kari K, Kerosuo E, Haapasalo M, Egelseer EM, Schocher I, Sleytr UB, Morelli L, Callegari ML, Nomellini JF, Bingle WH, Smit J, Leibovitz E, Lemaire M, Miras I, Salamitou S, Béguin P, Ohayon H, Gounon P, Matuschek M, Koval SF (1997). "Functions of S-layers". FEMS Microbiol Rev. 20 (1–2): 99–149. PMID 9276929. doi:10.1016/S0168-6445(97)00043-0.
- ↑ Kojima S, Blair DF (2004). "The bacterial flagellar motor: structure and function of a complex molecular machine". Int Rev Cytol. International Review of Cytology. 233: 93–134. ISBN 978-0-12-364637-8. PMID 15037363. doi:10.1016/S0074-7696(04)33003-2.
- ↑ Beachey EH (1981). "Bacterial adherence: adhesin-receptor interactions mediating the attachment of bacteria to mucosal surface". J Infect Dis. 143 (3): 325–45. PMID 7014727. doi:10.1093/infdis/143.3.325.
- ↑ Silverman PM (1997). "Towards a structural biology of bacterial conjugation". Mol Microbiol. 23 (3): 423–9. PMID 9044277. doi:10.1046/j.1365-2958.1997.2411604.x.
- ↑ Stokes RW, Norris-Jones R, Brooks DE, Beveridge TJ, Doxsee D, Thorson LM (2004). "The glycan-rich outer layer of the cell wall of Mycobacterium tuberculosis acts as an antiphagocytic capsule limiting the association of the bacterium with macrophages". Infect Immun. 72 (10): 5676–86. PMC 517526
 . PMID 15385466. doi:10.1128/IAI.72.10.5676-5686.2004.
. PMID 15385466. doi:10.1128/IAI.72.10.5676-5686.2004. - ↑ Daffé M, Etienne G (1999). "The capsule of Mycobacterium tuberculosis and its implications for pathogenicity". Tuber Lung Dis. 79 (3): 153–69. PMID 10656114. doi:10.1054/tuld.1998.0200.
- ↑ Finlay BB, Falkow S (1997). "Common themes in microbial pathogenicity revisited". Microbiology and Molecular Biology Reviews. 61 (2): 136–69. PMC 232605
 . PMID 9184008.
. PMID 9184008. - ↑ Nicholson WL, Munakata N, Horneck G, Melosh HJ, Setlow P (2000). "Resistance of Bacillus endospores to extreme terrestrial and extraterrestrial environments". Microbiology and Molecular Biology Reviews. 64 (3): 548–72. PMC 99004
 . PMID 10974126. doi:10.1128/MMBR.64.3.548-572.2000.
. PMID 10974126. doi:10.1128/MMBR.64.3.548-572.2000. - ↑ Siunov AV, Nikitin DV, Suzina NE, Dmitriev VV, Kuzmin NP, Duda VI (1999). "Phylogenetic status of Anaerobacter polyendosporus, an anaerobic, polysporogenic bacterium" (PDF). Int J Syst Bacteriol. 49 (3): 1119–24. PMID 10425769. doi:10.1099/00207713-49-3-1119.
- ↑ Nicholson WL, Fajardo-Cavazos P, Rebeil R, Slieman TA, Riesenman PJ, Law JF, Xue Y (2002). "Bacterial endospores and their significance in stress resistance". Antonie Van Leeuwenhoek. 81 (1–4): 27–32. PMID 12448702. doi:10.1023/A:1020561122764.
- ↑ Vreeland RH, Rosenzweig WD, Powers DW (2000). "Isolation of a 250 million-year-old halotolerant bacterium from a primary salt crystal". Nature. 407 (6806): 897–900. Bibcode:2000Natur.407..897V. PMID 11057666. doi:10.1038/35038060.
- ↑ Cano RJ, Borucki MK (1995). "Revival and identification of bacterial spores in 25- to 40-million-year-old Dominican amber". Science. 268 (5213): 1060–4. Bibcode:1995Sci...268.1060C. PMID 7538699. doi:10.1126/science.7538699.
- ↑ Nicholson WL, Schuerger AC, Setlow P (2005). "The solar UV environment and bacterial spore UV resistance: considerations for Earth-to-Mars transport by natural processes and human spaceflight". Mutat Res. 571 (1–2): 249–64. PMID 15748651. doi:10.1016/j.mrfmmm.2004.10.012.
- ↑ BBC Staff (23 August 2011). "Impacts 'more likely' to have spread life from Earth". BBC. Retrieved 24 August 2011.
- ↑ Hatheway CL (1990). "Toxigenic clostridia". Clinical Microbiology Reviews. 3 (1): 66–98. PMC 358141
 . PMID 2404569.
. PMID 2404569. - ↑ Nealson KH (1999). "Post-Viking microbiology: new approaches, new data, new insights". Origins of Life and Evolution of Biospheres. 29 (1): 73–93. PMID 11536899. doi:10.1023/A:1006515817767.
- ↑ Xu J (2006). "Microbial ecology in the age of genomics and metagenomics: concepts, tools, and recent advances". Mol Ecol. 15 (7): 1713–31. PMID 16689892. doi:10.1111/j.1365-294X.2006.02882.x.
- ↑ Zillig W (1991). "Comparative biochemistry of Archaea and Bacteria". Current Opinion in Genetics & Development. 1 (4): 544–51. PMID 1822288. doi:10.1016/S0959-437X(05)80206-0.
- ↑ Hellingwerf KJ, Crielaard W, Hoff WD, Matthijs HC, Mur LR, van Rotterdam BJ (1994). "Photobiology of bacteria". Antonie Van Leeuwenhoek. 65 (4): 331–47. PMID 7832590. doi:10.1007/BF00872217.
- ↑ Zumft WG (1 December 1997). "Cell biology and molecular basis of denitrification". Microbiol Mol Biol Rev. 61 (4): 533–616. PMC 232623
 . PMID 9409151.
. PMID 9409151. - ↑ Drake HL, Daniel SL, Küsel K, Matthies C, Kuhner C, Braus-Stromeyer S (1997). "Acetogenic bacteria: what are the in situ consequences of their diverse metabolic versatilities?". BioFactors. 6 (1): 13–24. PMID 9233536. doi:10.1002/biof.5520060103.
- ↑ Morel FM, Kraepiel AM, Amyot M (1998). "The chemical cycle and bioaccumulation of mercury". Annual Review of Ecology and Systematics. 29: 543–566. doi:10.1146/annurev.ecolsys.29.1.543.
- ↑ Dalton H (2005). "The Leeuwenhoek Lecture 2000 the natural and unnatural history of methane-oxidizing bacteria". Philosophical Transactions of the Royal Society B. 360 (1458): 1207–22. PMC 1569495
 . PMID 16147517. doi:10.1098/rstb.2005.1657.
. PMID 16147517. doi:10.1098/rstb.2005.1657. - ↑ Zehr JP, Jenkins BD, Short SM, Steward GF (2003). "Nitrogenase gene diversity and microbial community structure: a cross-system comparison". Environ Microbiol. 5 (7): 539–54. PMID 12823187. doi:10.1046/j.1462-2920.2003.00451.x.
- ↑ Lonhienne TG, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA (2010). "Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus". Proceedings of the National Academy of Sciences of the United States of America. 107 (29): 12883–12888. Bibcode:2010PNAS..10712883L. PMC 2919973
 . PMID 20566852. doi:10.1073/pnas.1001085107.
. PMID 20566852. doi:10.1073/pnas.1001085107. - ↑ Koch AL (2002). "Control of the bacterial cell cycle by cytoplasmic growth". Crit Rev Microbiol. 28 (1): 61–77. PMID 12003041. doi:10.1080/1040-840291046696.
- ↑ Eagon RG (1962). "Pseudomonas natriegens, a marine bacterium with a generation time of less than 10 minutes". Journal of Bacteriology. 83 (4): 736–7. PMC 279347
 . PMID 13888946.
. PMID 13888946. - ↑ Stewart EJ, Madden R, Paul G, Taddei F (2005). "Aging and death in an organism that reproduces by morphologically symmetric division". PLoS Biol. 3 (2): e45. PMC 546039
 . PMID 15685293. doi:10.1371/journal.pbio.0030045.
. PMID 15685293. doi:10.1371/journal.pbio.0030045. - 1 2 3 Thomson RB, Bertram H (2001). "Laboratory diagnosis of central nervous system infections". Infectious Disease Clinics of North America. 15 (4): 1047–71. PMID 11780267. doi:10.1016/S0891-5520(05)70186-0.
- ↑ Paerl HW, Fulton RS, Moisander PH, Dyble J (2001). "Harmful freshwater algal blooms, with an emphasis on cyanobacteria". ScientificWorldJournal. 1: 76–113. PMID 12805693. doi:10.1100/tsw.2001.16.
- ↑ Challis GL, Hopwood DA (2003). "Synergy and contingency as driving forces for the evolution of multiple secondary metabolite production by Streptomyces species". Proceedings of the National Academy of Sciences of the United States of America. 100 Suppl 2 (90002): 14555–61. Bibcode:2003PNAS..10014555C. PMC 304118
 . PMID 12970466. doi:10.1073/pnas.1934677100.
. PMID 12970466. doi:10.1073/pnas.1934677100. - ↑ Kooijman SA, Auger P, Poggiale JC, Kooi BW (2003). "Quantitative steps in symbiogenesis and the evolution of homeostasis". Biol Rev Camb Philos Soc. 78 (3): 435–63. PMID 14558592. doi:10.1017/S1464793102006127.
- ↑ Prats C, López D, Giró A, Ferrer J, Valls J (2006). "Individual-based modelling of bacterial cultures to study the microscopic causes of the lag phase". J Theor Biol. 241 (4): 939–53. PMID 16524598. doi:10.1016/j.jtbi.2006.01.029.
- ↑ Hecker M, Völker U (2001). "General stress response of Bacillus subtilis and other bacteria". Adv Microb Physiol. Advances in Microbial Physiology. 44: 35–91. ISBN 978-0-12-027744-5. PMID 11407115. doi:10.1016/S0065-2911(01)44011-2.
- ↑ "Genomes Online Database". Genomes Online Database. Joint Genome Institute. Retrieved 14 Sep 2016.
- ↑ Nakabachi A, Yamashita A, Toh H, Ishikawa H, Dunbar HE, Moran NA, Hattori M (2006). "The 160-kilobase genome of the bacterial endosymbiont Carsonella". Science. 314 (5797): 267. PMID 17038615. doi:10.1126/science.1134196.
- ↑ Pradella S, Hans A, Spröer C, Reichenbach H, Gerth K, Beyer S (2002). "Characterisation, genome size and genetic manipulation of the myxobacterium Sorangium cellulosum So ce56". Arch Microbiol. 178 (6): 484–92. PMID 12420170. doi:10.1007/s00203-002-0479-2.
- ↑ Belfort M, Reaban ME, Coetzee T, Dalgaard JZ (1 July 1995). "Prokaryotic introns and inteins: a panoply of form and function". J. Bacteriol. 177 (14): 3897–903. PMC 177115
 . PMID 7608058.
. PMID 7608058. - ↑ Hinnebusch J, Tilly K (1993). "Linear plasmids and chromosomes in bacteria". Mol Microbiol. 10 (5): 917–22. PMID 7934868. doi:10.1111/j.1365-2958.1993.tb00963.x.
- ↑ Fraser, Claire M.; Casjens, Sherwood; Huang, Wai Mun; Sutton, Granger G.; Clayton, Rebecca; Lathigra, Raju; White, Owen; Ketchum, Karen A.; Dodson, Robert (1997-12-11). "Genomic sequence of a Lyme disease spirochaete, Borrelia burgdorferi". Nature. 390 (6660): 580–586. ISSN 0028-0836. PMID 9403685. doi:10.1038/37551.
- ↑ The University of Waikato (March 25, 2014). "Bacterial DNA – the role of plasmids". Themes — Bacteria in biotech. Biotechnology Learning Hub. Retrieved 2014-09-03.
- ↑ Denamur E, Matic I (2006). "Evolution of mutation rates in bacteria". Mol Microbiol. 60 (4): 820–7. PMID 16677295. doi:10.1111/j.1365-2958.2006.05150.x.
- ↑ Wright BE (2004). "Stress-directed adaptive mutations and evolution". Mol Microbiol. 52 (3): 643–50. PMID 15101972. doi:10.1111/j.1365-2958.2004.04012.x.
- ↑ Chen I, Dubnau D (2004). "DNA uptake during bacterial transformation". Nature Reviews Microbiology. 2 (3): 241–9. PMID 15083159. doi:10.1038/nrmicro844.
- ↑ Solomon JM, Grossman AD (1996). "Who's competent and when: regulation of natural genetic competence in bacteria". Trends Genet. 12 (4): 150–5. PMID 8901420. doi:10.1016/0168-9525(96)10014-7.
- ↑ Akamatsu T, Taguchi H (2001). "Incorporation of the whole chromosomal DNA in protoplast lysates into competent cells of Bacillus subtilis". Biosci. Biotechnol. Biochem. 65 (4): 823–9. PMID 11388459. doi:10.1271/bbb.65.823.
- ↑ Saito Y, Taguchi H, Akamatsu T (2006). "Fate of transforming bacterial genome following incorporation into competent cells of Bacillus subtilis: a continuous length of incorporated DNA". J. Biosci. Bioeng. 101 (3): 257–62. PMID 16716928. doi:10.1263/jbb.101.257.
- ↑ Johnsborg O, Eldholm V, Håvarstein LS (2007). "Natural genetic transformation: prevalence, mechanisms and function". Res. Microbiol. 158 (10): 767–78. PMID 17997281. doi:10.1016/j.resmic.2007.09.004.
- ↑ Bernstein H, Bernstein C, Michod RE (2012). "DNA repair as the primary adaptive function of sex in bacteria and eukaryotes". Chapter 1: pp. 1–49 in: DNA Repair: New Research, Sakura Kimura and Sora Shimizu (eds.). Nova Sci. Publ., Hauppauge, N.Y. ISBN 978-1-62100-808-8.
- ↑ Michod RE, Bernstein H, Nedelcu AM (2008). "Adaptive value of sex in microbial pathogens" (PDF). Infect. Genet. Evol. 8 (3): 267–85. PMID 18295550. doi:10.1016/j.meegid.2008.01.002.
- ↑ Davison J (1999). "Genetic exchange between bacteria in the environment". Plasmid. 42 (2): 73–91. PMID 10489325. doi:10.1006/plas.1999.1421.
- ↑ Hastings PJ, Rosenberg SM, Slack A (2004). "Antibiotic-induced lateral transfer of antibiotic resistance". Trends Microbiol. 12 (9): 401–4. PMID 15337159. doi:10.1016/j.tim.2004.07.003.
- ↑ Brüssow H, Canchaya C, Hardt WD (2004). "Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion". Microbiology and Molecular Biology Reviews. 68 (3): 560–602. PMC 515249
 . PMID 15353570. doi:10.1128/MMBR.68.3.560-602.2004.
. PMID 15353570. doi:10.1128/MMBR.68.3.560-602.2004. - ↑ Bickle TA, Krüger DH (1993). "Biology of DNA restriction". Microbiol. Rev. 57 (2): 434–50. PMC 372918
 . PMID 8336674.
. PMID 8336674. - ↑ Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007). "CRISPR provides acquired resistance against viruses in prokaryotes". Science. 315 (5819): 1709–12. Bibcode:2007Sci...315.1709B. PMID 17379808. doi:10.1126/science.1138140.
- ↑ Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, van der Oost J (2008). "Small CRISPR RNAs guide antiviral defense in prokaryotes". Science. 321 (5891): 960–4. Bibcode:2008Sci...321..960B. PMID 18703739. doi:10.1126/science.1159689.
- ↑ Dusenbery, David B. (1996). Life at Small Scale. Scientific American Library. ISBN 0-7167-5060-0.
- 1 2 Shapiro JA (1998). "Thinking about bacterial populations as multicellular organisms" (PDF). Annu. Rev. Microbiol. 52: 81–104. PMID 9891794. doi:10.1146/annurev.micro.52.1.81. Archived from the original (PDF) on 17 July 2011.
- 1 2 Costerton JW, Lewandowski Z, Caldwell DE, Korber DR, Lappin-Scott HM (1995). "Microbial biofilms". Annu. Rev. Microbiol. 49: 711–45. PMID 8561477. doi:10.1146/annurev.mi.49.100195.003431.
- ↑ Miller MB, Bassler BL (2001). "Quorum sensing in bacteria". Annu. Rev. Microbiol. 55: 165–99. PMID 11544353. doi:10.1146/annurev.micro.55.1.165.
- 1 2 3 Bardy S, Ng S, Jarrell K (2003). "Prokaryotic motility structures". Microbiology. 149 (Pt 2): 295–304. PMID 12624192. doi:10.1099/mic.0.25948-0.
- ↑ Dusenbery, David B. (2009). Living at Micro Scale, p. 136. Harvard University Press, Cambridge, Mass. ISBN 978-0-674-03116-6.
- ↑ Merz A, So M, Sheetz M (2000). "Pilus retraction powers bacterial twitching motility". Nature. 407 (6800): 98–102. Bibcode:2000Natur.407...98M. PMID 10993081. doi:10.1038/35024105.
- ↑ "A re-examination of twitching motility in Pseudomonas aeruginosa" – Semmler, Whitchurch & Mattick (1999)
- ↑ Dusenbery, David B. (2009). Living at Micro Scale, Chapter 13. Harvard University Press, Cambridge, Mass. ISBN 978-0-674-03116-6.
- ↑ Macnab RM (1 December 1999). "The bacterial flagellum: reversible rotary propellor and type III export apparatus". J. Bacteriol. 181 (23): 7149–53. PMC 103673
 . PMID 10572114.
. PMID 10572114. - ↑ Wu M, Roberts J, Kim S, Koch D, DeLisa M (2006). "Collective bacterial dynamics revealed using a three-dimensional population-scale defocused particle tracking technique". Appl Environ Microbiol. 72 (7): 4987–94. PMC 1489374
 . PMID 16820497. doi:10.1128/AEM.00158-06.
. PMID 16820497. doi:10.1128/AEM.00158-06. - ↑ Lux R, Shi W (2004). "Chemotaxis-guided movements in bacteria". Crit Rev Oral Biol Med. 15 (4): 207–20. PMID 15284186. doi:10.1177/154411130401500404.
- ↑ Schweinitzer T, Josenhans C (2010). "Bacterial energy taxis: a global strategy?". Arch Microbiol. 192 (7): 507–20. PMC 2886117
 . PMID 20411245. doi:10.1007/s00203-010-0575-7.
. PMID 20411245. doi:10.1007/s00203-010-0575-7. - ↑ Frankel R, Bazylinski D, Johnson M, Taylor B (1997). "Magneto-aerotaxis in marine coccoid bacteria". Biophys J. 73 (2): 994–1000. Bibcode:1997BpJ....73..994F. PMC 1180996
 . PMID 9251816. doi:10.1016/S0006-3495(97)78132-3.
. PMID 9251816. doi:10.1016/S0006-3495(97)78132-3. - ↑ Goldberg MB (2001). "Actin-based motility of intracellular microbial pathogens". Microbiol Mol Biol Rev. 65 (4): 595–626, table of contents. PMC 99042
 . PMID 11729265. doi:10.1128/MMBR.65.4.595-626.2001.
. PMID 11729265. doi:10.1128/MMBR.65.4.595-626.2001. - ↑ Boucher Y, Douady CJ, Papke RT, Walsh DA, Boudreau ME, Nesbo CL, Case RJ, Doolittle WF (2003). "Lateral gene transfer and the origins of prokaryotic groups". Annu Rev Genet. 37: 283–328. PMID 14616063. doi:10.1146/annurev.genet.37.050503.084247.
- ↑ Olsen GJ, Woese CR, Overbeek R (1994). "The winds of (evolutionary) change: breathing new life into microbiology". Journal of Bacteriology. 176 (1): 1–6. PMC 205007
 . PMID 8282683. doi:10.2172/205047.
. PMID 8282683. doi:10.2172/205047. - ↑ "IJSEM Home". Ijs.sgmjournals.org. 28 October 2011. Retrieved 4 November 2011.
- ↑ "Bergey's Manual Trust". Bergeys.org. Retrieved 4 November 2011.
- ↑ Gupta R (2000). "The natural evolutionary relationships among prokaryotes". Crit Rev Microbiol. 26 (2): 111–31. PMID 10890353. doi:10.1080/10408410091154219.
- ↑ Doolittle RF (2005). "Evolutionary aspects of whole-genome biology". Current Opinion in Structural Biology. 15 (3): 248–53. PMID 15963888. doi:10.1016/j.sbi.2005.04.001.
- ↑ Cavalier-Smith T (2002). "The neomuran origin of archaebacteria, the negibacterial root of the universal tree and bacterial megaclassification". Int J Syst Evol Microbiol. 52 (Pt 1): 7–76. PMID 11837318. doi:10.1099/00207713-52-1-7.
- ↑ Woods GL, Walker DH (1996). "Detection of infection or infectious agents by use of cytologic and histologic stains". Clinical Microbiology Reviews. 9 (3): 382–404. PMC 172900
 . PMID 8809467.
. PMID 8809467. - ↑ Weinstein M (1994). "Clinical importance of blood cultures". Clin Lab Med. 14 (1): 9–16. PMID 8181237.
- ↑ Louie M, Louie L, Simor AE (8 August 2000). "The role of DNA amplification technology in the diagnosis of infectious diseases". CMAJ. 163 (3): 301–9. PMC 80298
 . PMID 10951731.
. PMID 10951731. - ↑ Oliver J (2005). "The viable but nonculturable state in bacteria". J Microbiol. 43 Spec No: 93–100. PMID 15765062. Archived from the original on 28 September 2007.
- ↑ Euzéby JP (8 December 2011). "Number of published names". List of Prokaryotic names with Standing in Nomenclature. Archived from the original on 19 January 2012. Retrieved 10 December 2011.
- ↑ Curtis TP, Sloan WT, Scannell JW (2002). "Estimating prokaryotic diversity and its limits". Proceedings of the National Academy of Sciences of the United States of America. 99 (16): 10494–9. Bibcode:2002PNAS...9910494C. PMC 124953
 . PMID 12097644. doi:10.1073/pnas.142680199.
. PMID 12097644. doi:10.1073/pnas.142680199. - ↑ Schloss PD, Handelsman J (2004). "Status of the microbial census". Microbiology and Molecular Biology Reviews. 68 (4): 686–91. PMC 539005
 . PMID 15590780. doi:10.1128/MMBR.68.4.686-691.2004.
. PMID 15590780. doi:10.1128/MMBR.68.4.686-691.2004. - ↑ Fisher B, Harvey RP, Champe PC (2007). Lippincott's Illustrated Reviews: Microbiology (Lippincott's Illustrated Reviews Series). Hagerstwon, MD: Lippincott Williams & Wilkins. pp. Chapter 33, pages 367–392. ISBN 0-7817-8215-5.
- ↑ LEF.org > Bacterial Infections Updated: 19 January 2006. Retrieved on 11 April 2009
- ↑ Martin MO (2002). "Predatory prokaryotes: an emerging research opportunity". Journal of Microbiology and Biotechnology. 4 (5): 467–77. PMID 12432957.
- ↑ Velicer GJ, Stredwick KL (2002). "Experimental social evolution with Myxococcus xanthus". Antonie Van Leeuwenhoek. 81 (1–4): 155–64. PMID 12448714. doi:10.1023/A:1020546130033.
- ↑ Gromov BV (1972). "Electron Microscope Study of Parasitism by Bdellovibrio Chorellavorus Bacteria on Cells of the Green Alga Chorella Vulgaris". Tsitologiya. 14 (2): 256–60.
- ↑ Guerrero R, Pedros-Alio C, Esteve I, Mas J, Chase D, Margulis L (April 1986). "Predatory prokaryotes: predation and primary consumption evolved in bacteria". Proceedings of the National Academy of Sciences of the United States of America. 83 (7): 2138–42. Bibcode:1986PNAS...83.2138G. PMC 323246
 . PMID 11542073. doi:10.1073/pnas.83.7.2138.
. PMID 11542073. doi:10.1073/pnas.83.7.2138. - ↑ Velicer GJ, Mendes-Soares H (2009). "Bacterial predators". Current Biology. 19 (2): R55–6. PMID 19174136. doi:10.1016/j.cub.2008.10.043.
- ↑ Stams AJ, de Bok FA, Plugge CM, van Eekert MH, Dolfing J, Schraa G (2006). "Exocellular electron transfer in anaerobic microbial communities". Environ Microbiol. 8 (3): 371–82. PMID 16478444. doi:10.1111/j.1462-2920.2006.00989.x.
- ↑ Barea JM, Pozo MJ, Azcón R, Azcón-Aguilar C (2005). "Microbial co-operation in the rhizosphere". J Exp Bot. 56 (417): 1761–78. PMID 15911555. doi:10.1093/jxb/eri197.
- ↑ O'Hara AM, Shanahan F (2006). "The gut flora as a forgotten organ". EMBO Reports. 7 (7): 688–93. PMC 1500832
 . PMID 16819463. doi:10.1038/sj.embor.7400731.
. PMID 16819463. doi:10.1038/sj.embor.7400731. - ↑ Zoetendal EG, Vaughan EE, de Vos WM (2006). "A microbial world within us". Mol Microbiol. 59 (6): 1639–50. PMID 16553872. doi:10.1111/j.1365-2958.2006.05056.x.
- ↑ Gorbach SL (1990). "Lactic acid bacteria and human health". Annals of Medicine. 22 (1): 37–41. PMID 2109988. doi:10.3109/07853899009147239.
- ↑ Salminen SJ, Gueimonde M, Isolauri E (1 May 2005). "Probiotics that modify disease risk". J Nutr. 135 (5): 1294–8. PMID 15867327.
- ↑ Fish DN (2002). "Optimal antimicrobial therapy for sepsis". Am J Health Syst Pharm. 59 Suppl 1: S13–9. PMID 11885408.
- ↑ Belland RJ, Ouellette SP, Gieffers J, Byrne GI (2004). "Chlamydia pneumoniae and atherosclerosis". Cell Microbiol. 6 (2): 117–27. PMID 14706098. doi:10.1046/j.1462-5822.2003.00352.x.
- ↑ Heise ER (1982). "Diseases associated with immunosuppression". Environmental Health Perspectives. 43: 9–19. JSTOR 3429162. PMC 1568899
 . PMID 7037390. doi:10.2307/3429162.
. PMID 7037390. doi:10.2307/3429162. - ↑ Saiman L (2004). "Microbiology of early CF lung disease". Paediatric Respiratory Reviews. 5 Suppl A: S367–9. PMID 14980298. doi:10.1016/S1526-0542(04)90065-6.
- ↑ Yonath A, Bashan A (2004). "Ribosomal crystallography: initiation, peptide bond formation, and amino acid polymerization are hampered by antibiotics". Annu Rev Microbiol. 58: 233–51. PMID 15487937. doi:10.1146/annurev.micro.58.030603.123822.
- ↑ Khachatourians GG (1998). "Agricultural use of antibiotics and the evolution and transfer of antibiotic-resistant bacteria". CMAJ. 159 (9): 1129–36. PMC 1229782
 . PMID 9835883.
. PMID 9835883. - ↑ Johnson ME, Lucey JA (2006). "Major technological advances and trends in cheese". J Dairy Sci. 89 (4): 1174–8. PMID 16537950. doi:10.3168/jds.S0022-0302(06)72186-5.
- ↑ Hagedorn S, Kaphammer B (1994). "Microbial biocatalysis in the generation of flavor and fragrance chemicals". Annu. Rev. Microbiol. 48: 773–800. PMID 7826026. doi:10.1146/annurev.mi.48.100194.004013.
- ↑ Cohen Y (2002). "Bioremediation of oil by marine microbial mats". Int Microbiol. 5 (4): 189–93. PMID 12497184. doi:10.1007/s10123-002-0089-5.
- ↑ Neves LC, Miyamura TT, Moraes DA, Penna TC, Converti A (2006). "Biofiltration methods for the removal of phenolic residues". Appl. Biochem. Biotechnol. 129–132: 130–52. PMID 16915636. doi:10.1385/ABAB:129:1:130.
- ↑ Liese A, Filho MV (1999). "Production of fine chemicals using biocatalysis". Current Opinion in Biotechnology. 10 (6): 595–603. PMID 10600695. doi:10.1016/S0958-1669(99)00040-3.
- ↑ Aronson AI, Shai Y (2001). "Why Bacillus thuringiensis insecticidal toxins are so effective: unique features of their mode of action". FEMS Microbiol. Lett. 195 (1): 1–8. PMID 11166987. doi:10.1111/j.1574-6968.2001.tb10489.x.
- ↑ Bozsik A (2006). "Susceptibility of adult Coccinella septempunctata (Coleoptera: Coccinellidae) to insecticides with different modes of action". Pest Manag Sci. 62 (7): 651–4. PMID 16649191. doi:10.1002/ps.1221.
- ↑ Chattopadhyay A, Bhatnagar NB, Bhatnagar R (2004). "Bacterial insecticidal toxins". Crit Rev Microbiol. 30 (1): 33–54. PMID 15116762. doi:10.1080/10408410490270712.
- ↑ Serres MH, Gopal S, Nahum LA, Liang P, Gaasterland T, Riley M (2001). "A functional update of the Escherichia coli K-12 genome". Genome Biology. 2 (9): research0035.1–research0035.7. PMC 56896
 . PMID 11574054. doi:10.1186/gb-2001-2-9-research0035.
. PMID 11574054. doi:10.1186/gb-2001-2-9-research0035. - ↑ Almaas E, Kovács B, Vicsek T, Oltvai ZN, Barabási AL (2004). "Global organization of metabolic fluxes in the bacterium Escherichia coli". Nature. 427 (6977): 839–43. Bibcode:2004Natur.427..839A. PMID 14985762. arXiv:q-bio/0403001
 . doi:10.1038/nature02289.
. doi:10.1038/nature02289. - ↑ Reed JL, Vo TD, Schilling CH, Palsson BO (2003). "An expanded genome-scale model of Escherichia coli K-12 (iJR904 GSM/GPR)". Genome Biol. 4 (9): R54. PMC 193654
 . PMID 12952533. doi:10.1186/gb-2003-4-9-r54.
. PMID 12952533. doi:10.1186/gb-2003-4-9-r54. - ↑ Walsh G (2005). "Therapeutic insulins and their large-scale manufacture". Appl Microbiol Biotechnol. 67 (2): 151–9. PMID 15580495. doi:10.1007/s00253-004-1809-x.
- ↑ Graumann K, Premstaller A (2006). "Manufacturing of recombinant therapeutic proteins in microbial systems". Biotechnol J. 1 (2): 164–86. PMID 16892246. doi:10.1002/biot.200500051.
- ↑ Porter JR (1976). "Antony van Leeuwenhoek: tercentenary of his discovery of bacteria". Bacteriological Reviews. 40 (2): 260–9. PMC 413956
 . PMID 786250.
. PMID 786250. - ↑ van Leeuwenhoek A (1684). "An abstract of a letter from Mr. Anthony Leevvenhoek at Delft, dated Sep. 17, 1683, Containing Some Microscopical Observations, about Animals in the Scurf of the Teeth, the Substance Call'd Worms in the Nose, the Cuticula Consisting of Scales". Philosophical Transactions (1683–1775). 14 (155–166): 568–574. doi:10.1098/rstl.1684.0030.
- ↑ van Leeuwenhoek A (1700). "Part of a Letter from Mr Antony van Leeuwenhoek, concerning the Worms in Sheeps Livers, Gnats, and Animalcula in the Excrements of Frogs". Philosophical Transactions (1683–1775). 22 (260–276): 509–518. doi:10.1098/rstl.1700.0013.
- ↑ van Leeuwenhoek A (1702). "Part of a Letter from Mr Antony van Leeuwenhoek, F. R. S. concerning Green Weeds Growing in Water, and Some Animalcula Found about Them". Philosophical Transactions (1683–1775). 23 (277–288): 1304–11. doi:10.1098/rstl.1702.0042.
- ↑ Asimov, Isaac (1982), Asimov's Biographical Encyclopedia of Science and Technology, 2nd edition, Garden City, New York: Doubleday and Company, pg 143.
- ↑ Ehrenberg's Symbolae Physioe. Animalia evertebrata. Decas prima. Berlin, 1828.
- ↑ Breed RS, Conn HJ (1936). "The Status of the Generic Term Bacterium Ehrenberg 1828". Journal of Bacteriology. 31 (5): 517–518. PMC 543738
 . PMID 16559906.
. PMID 16559906. - ↑ EHRENBERG (C.G.): Dritter Beitrag zur Erkenntniss grosser Organisation in der Richtung des kleinsten Raumes. Physikalische Abhandlungen der Koeniglichen Akademie der Wissenschaften zu Berlin aus den Jahren 1833–1835, 1835, pp. 143–336.
- ↑ "Pasteur's Papers on the Germ Theory". LSU Law Center's Medical and Public Health Law Site, Historic Public Health Articles. Archived from the original on 18 December 2006. Retrieved 23 November 2006.
- ↑ "The Nobel Prize in Physiology or Medicine 1905". Nobelprize.org. Archived from the original on 10 December 2006. Retrieved 22 November 2006.
- ↑ O'Brien SJ, Goedert JJ (1996). "HIV causes AIDS: Koch's postulates fulfilled". Current Opinion in Immunology. 8 (5): 613–8. PMID 8902385. doi:10.1016/S0952-7915(96)80075-6.
- ↑ Thurston AJ (2000). "Of blood, inflammation and gunshot wounds: the history of the control of sepsis". Aust N Z J Surg. 70 (12): 855–61. PMID 11167573. doi:10.1046/j.1440-1622.2000.01983.x.
- ↑ Schwartz RS (2004). "Paul Ehrlich's magic bullets". N Engl J Med. 350 (11): 1079–80. PMID 15014180. doi:10.1056/NEJMp048021.
- ↑ "Biography of Paul Ehrlich". Nobelprize.org. Archived from the original on 28 November 2006. Retrieved 26 November 2006.
Further reading
| Library resources about Bacteria |
- Alcamo IE (2001). Fundamentals of microbiology. Boston: Jones and Bartlett. ISBN 0-7637-1067-9.
- Atlas RM (1995). Principles of microbiology. St. Louis: Mosby. ISBN 0-8016-7790-4.
- Martinko JM, Madigan MT (2005). Brock Biology of Microorganisms (11th ed.). Englewood Cliffs, N.J: Prentice Hall. ISBN 0-13-144329-1.
- Holt JC, Bergey DH (1994). Bergey's manual of determinative bacteriology (9th ed.). Baltimore: Williams & Wilkins. ISBN 0-683-00603-7.
- Hugenholtz P, Goebel BM, Pace NR (15 September 1998). "Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity". J Bacteriol. 180 (18): 4765–74. PMC 107498
 . PMID 9733676.
. PMID 9733676. - Funke BR, Tortora GJ, Case CL (2004). Microbiology: an introduction (8th ed.). San Francisco: Benjamin Cummings. ISBN 0-8053-7614-3.
- Ogunseitan OA (2005). Microbial Diversity: Form and Function in Prokaryotes. Wiley-Blackwell. ISBN 978-1-4051-4448-3.
- Shively JM (2006). Complex Intracellular Structures in Prokaryotes (Microbiology Monographs). Berlin: Springer. ISBN 3-540-32524-7.
External links
- MicrobeWiki, an extensive wiki about bacteria and viruses
- Bacteria that affect crops and other plants
- Bacterial Nomenclature Up-To-Date from DSMZ
- Genera of the domain Bacteria—list of Prokaryotic names with Standing in Nomenclature
- The largest bacteria
- Tree of Life: Eubacteria
- Videos of bacteria swimming and tumbling, use of optical tweezers and other videos.
- Planet of the Bacteria by Stephen Jay Gould
- On-line text book on bacteriology
- Animated guide to bacterial cell structure.
- Bacteria Make Major Evolutionary Shift in the Lab
- Online collaboration for bacterial taxonomy.
- PATRIC, a Bioinformatics Resource Center for bacterial pathogens, funded by NIAID
- Bacterial Chemotaxis Interactive Simulator—A web-app that uses several simple algorithms to simulate bacterial chemotaxis.
- Cell-Cell Communication in Bacteria on-line lecture by Bonnie Bassler, and TED: Discovering bacteria's amazing communication system
- Sulfur-cycling fossil bacteria from the 1.8-Ga Duck Creek Formation provide promising evidence of evolution's null hypothesis, Proceedings of the National Academy of Sciences of the United States of America. Summarised in: Scientists discover bacteria that haven't evolved in more than 2 billion years, LiveScience and BusinessInsider