Silencer (DNA)
In genetics, a silencer is a DNA sequence capable of binding transcription regulation factors, called repressors. DNA contains genes and provides the template to produce messenger RNA (mRNA). That mRNA is then translated into proteins that activate or inactivate gene expression in cells. When a repressor protein binds to the silencer region of DNA, RNA polymerase—the enzyme that transcribes DNA into RNA—is prevented from binding to the promoter region. With the transcription of DNA into RNA blocked, the translation of RNA into proteins is impossible. Thus, silencers prevent genes from being expressed as proteins.
RNA polymerase, a DNA-dependent enzyme, transcribes the DNA sequences, called nucleotides, in the 3' to 5' direction while the complementary RNA is synthesized in the 5' to 3' direction. RNA is similar to DNA, except that RNA contains uracil, instead of thymine, which forms a base pair with adenine. An important region for the activity of gene repression and expression found in RNA is the 3' untranslated region. This is a region on the 3' terminus of RNA that will not be translated to protein but includes many regulatory regions.
Not much is yet known about silencers but scientists continue to study in hopes to classify more types, locations in the genome, and diseases associated with silencers.

Functionality
Locations within the genome
A silencer is a sequence-specific element that induces a negative effect on the transcription of its particular gene. There are many positions in which a silencer element can be located in DNA. The most common position is found upstream of the target gene where it can help repress the transcription of the gene.[1] This distance can vary greatly between approximately -20 bp to -2000 bp upstream of a gene. Certain silencers can be found downstream of a promoter located within the intron or exon of the gene itself. Silencers have also been found within the 3 prime untranslated region (3' UTR) of mRNA.[2]
Types
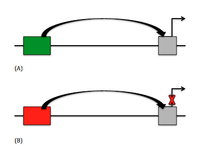
Currently, there are two main types of silencers in DNA, which are the classical silencer element and the non-classical negative regulatory element (NRE). In classical silencers, the gene is actively repressed by the silencer element, mostly by interfering with general transcription factor (GTF) assembly.[2] NREs passively repress the gene, usually by inhibiting other elements that are upstream of the gene. Of the NREs, there are certain silencers that are orientation-dependent meaning that the binding factor binds in a particular direction relative to other sequences. Promoter-dependent silencers are understood to be silencer elements because they are position and orientation-dependent but must also use a promoter-specific factor.[2] There has been a recent discovery of Polycomb-group Response Elements (PREs), which can allow and inhibit repression depending on the protein bound to it, and the presence of non-coding transcription.[1]
Mechanisms
For classical silencers, the signaling pathway is relatively simple. Since repression is active, silencer elements target the assembly of GTFs, necessary for transcription of the gene. These silencer elements are mostly located upstream of the gene and can vary between short and long distances. For long-range silencers, it has been observed that the DNA will form a loop in order to bring the silencer closer to the promoter and loop out the interfering DNA.[1] Silencers also target helicase sites in the DNA that are rich in adenine and thymine (AT) and prone to unwinding the DNA, allowing room to initiate transcription. The inhibited helicase activity leads to the inhibition of transcription. This is commonly seen in the human thyrotropin-β gene promoter. NREs can induce a bend in the promoter region to block interactions, as seen when an NRE binds to Yin-Yang 1 (YY1),[2] and flank regulatory signals or promoter regions as well. When the silencer region is located within an intron, there can be two types of repressions. First, there can be a physical blockage of a splice site. Second, there can be a bend in the DNA that will inhibit RNA processing.[2]
When located in the exon or the untranslated region, the silencer will mainly be classical or position-dependent. However, these silencers can carry out their activity prior to transcription.[2] Most silencers are constitutively expressed in organisms, only allowing activation of a gene by either inhibiting the silencer or by activating an enhancer region. The best example of this is the Neuronal-Restrictive Silencer Factor (NRSF) that is produced by the REST gene. The REST gene produces NRSF in order to repress the transcription of neuronal genes that are essential for localization of neuronal tissue. When a silencer represses REST, NRSF is also inhibited, allowing for the transcription of neuronal genes.[2]
Similarities with enhancers
Another regulatory element located upstream of the gene is an enhancer. Enhancers function as a "turn on" switch in gene expression and will activate the promoter region of a particular gene while silencers act as the "turn off" switch. Though these two regulatory elements work against each other, both sequence types affect the promoter region in very similar ways.[1] Because silencers have not been thoroughly identified and analyzed, the extensive research on enhancers has aided biologists in understanding the mechanics of the silencer. Enhancers can be found in many of the same areas that silencers are found, such as upstream of the promoter by many kilobase pairs, or even downstream within the intron of the gene.[1] DNA looping is also a model function used by enhancers in order to shorten the proximity of the promoter to the enhancer. Enhancers also function with transcription factors in order to initiate expression, much like silencers can with repressors.[1]
In prokaryotes and eukaryotes
Prokaryotes

There are several differences in the regulation of metabolic control in eukaryotes and in prokaryotes. Prokaryotes vary the numbers of specific enzymes made in their cells in order to regulate gene expression, which is slow metabolic control, and also regulate enzymatic pathways through mechanisms such as feedback inhibition and allosteric regulation, which is rapid metabolic control.[3] The genes of prokaryotes are grouped together based on similar functions into units called operons which consist of a promoter and an operator. The operator is the binding site for the repressor and thus has a function equivalent to the silencer region in Eukaryotic DNA. When a repressor protein is bound to the operator, RNA polymerase cannot bind to the promoter to initiate the transcription of the operon.
Repression of the lac operon
The lac operon in the prokaryote E. coli consists of genes that produce enzymes to break down lactose. Its operon is an example of a prokaryotic silencer. The three functional genes in this operon are lacZ, lacY, and lacA.[3] The repressor gene, lacI, will produce the repressor protein LacI which is under allosteric regulation. These genes are activated by the presence of lactose in the cell which acts as an effector molecule that binds to LacI. When the repressor is bound to lactose, it will not bind to the operator, which allows RNA polymerase to bind to the promoter to initiate transcription of the operon. When the repressor's allosteric site is not bound to lactose, its active site will bind to the operator to prevent RNA polymerase from transcribing the genes of the lac operon.
Eukaryotes
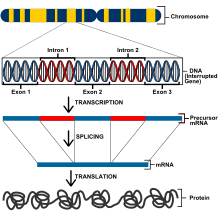
Eukaryotes have a much larger genome and thus have different methods of gene regulation than in prokaryotes. All cells in a eukaryotic organism have the same DNA but are specified through differential gene expression, a phenomenon known as genetic totipotency.[4] However, in order for a cell to express the genes for proper functioning, the genes must be closely regulated to express the correct properties. Genes in eukaryotes are controlled on the transcriptional, post-transcriptional, translational, and post-translational levels.[5] On the transcriptional level, gene expression is regulated by altering transcription rates. Genes that encode proteins include exons which will encode the polypeptides, introns that are removed from mRNA before the translation of proteins, a transcriptional start site in which RNA polymerase binds, and a promoter.[6]
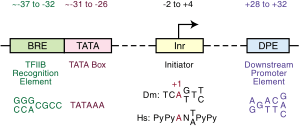
Repression of the TATA box
Eukaryotic genes contain an upstream promoter and a core promoter also referred to as a basal promoter. A common basal promoter is the TATAAAAAA sequence known as the TATA box. The TATA box is a complex with several different proteins including transcription factor II D (TFIID) which includes the TATA-binding protein (TBP) that binds to the TATA box along with 13 other proteins that bind to TBP. The TATA box binding proteins also include the transcription factor II B (TFIIB) which binds to both DNA and RNA polymerases.[6]
Silencers in eukaryotes control gene expression on a transcriptional level in which the mRNA is not transcribed. These DNA sequences may act as either silencers or enhancers based on the transcription factor that binds to the sequence and binding of this sequence will prevent promoters such as the TATA box from binding to RNA polymerase.[4] A repressor protein may have regions that bind to the DNA sequence as well as regions that bind to the transcription factors assembled at the promoter of the gene which would create a chromosome looping mechanism.[6] Looping brings silencers in close proximity to the promoters to ensure that groups of proteins needed for optimal gene expression will work together.
Mutated silencers, hereditary diseases, and their effects
Genetic mutations occur when nucleotide sequences in an organism are altered. These mutations lead to not only observable phenotypic influences in an individual, but also alterations that are undetectable phenotypically. The sources for these mutations can be errors during replication, spontaneous mutations, and chemical and physical mutagens (UV and ionizing radiation, heat).[7] Silencers, being encoded in the genome, are susceptible to such alterations which, in many cases, can lead to severe phenotypical and functional abnormalities. In general terms, mutations in silencer elements or regions could lead to either the inhibition of the silencer’s action or to the persisting repression of a necessary gene. This can then lead to the expression or suppression of an undesired phenotype which may affect the normal functionality of certain systems in the organism. Among the many silencer elements and proteins, REST/NSRF is an important silencer factor that has a variety of impacts, not only in neural aspects of development. In fact, in many cases, REST/NSRF acts in conjunction with RE-1/NRSE to repress and influence non-neuronal cells.[8] Its effects range from frogs (Xenopus laevis) to humans, with innumerous effects in phenotype and also in development. In Xenopus laevis, REST/NRSF malfunction or damage has been associated to abnormal ectodermal patterning during development and significant consequences in neural tube, cranial ganglia, and eye development.[9] In humans, a deficiency in the REST/NSRF silencer element has been correlated to Huntington's disease due to the decrease in the transcription of BDNF.
Furthermore, ongoing studies indicate that NRSE is involved in the regulation of the ANP gene, which when over expressed, can lead to ventricular hypertrophy.[10] Mutations in the Polycomb-group (PcG) complexes also presented significant modifications in physiological systems of organisms. Hence, modification in silencer elements and sequences can result in either devastating or unnoticeable changes.

REST/NRSF in Xenopus laevis
The effects and influences of RE1/NRSE and REST/NRSF are significant in non-neuronal cells that require the repression or silencing of neuronal genes. These silencer elements also regulate the expression of genes that do not induce neuron-specific proteins and studies have shown the extensive impact these factors have in cellular processes. In Xenopus laevis, RE1/NRSE and REST/NRSF dysfunction or mutation demonstrated significant impact on neural tube, cranial ganglia, and eye development.[9] All of these alterations can be traced to an improper patterning of the ectoderm during Xenopus development. Thus, a mutation or alteration in either the silencing region RE1/NRSE or silencer REST/NRSF factor can disrupt the proper differentiation and specification of the neuroepithelial domain and also hinder the formation of skin or ectoderm.[9] The lack of these factors result in a decreased production of bone morphogenetic protein (BMP), which translates into a deficient development of the neural crest.[9] Hence, the effects of NRSE and NRSF are of fundamental importance for neurogenesis of the developing embryo, and also in the early stages of ectodermal patterning. Ultimately, inadequate functioning of these factors can result in aberrant neural tube, cranial ganglia, and eye development in Xenopus.
REST/NSRF and Huntington's disease
Huntington's disease (HD) is an inherited neurodegenerative disorder, with symptoms emerging during an individual’s mid-adulthood. The most noticeable symptoms of this progressive disease are cognitive and motor impairments, as well as behavioral alterations.[11] These impairments can develop into dementia, chorea, and eventually death. At the molecular level, HD results from a mutation in the huntingtin protein (Htt). More specifically, there is an abnormal repetition of a CAG sequence towards the 5’-end of the gene, which then leads to the development of a toxic polyglutamine (polyQ) stretch in the protein. The mutated Htt protein affects an individual’s proper neural functions by inhibiting the action of REST/NRSF.
REST/NRSF is an important silencer element that binds to regulatory regions to control the expression of certain proteins involved in neural functions. The mechanistic actions of huntingtin are still not fully understood, but a correlation between Htt and REST/NRSF exists in HD development. By attaching to the REST/NRSF, the mutated huntingtin protein inhibits the action of the silencer element, and retains it in the cytosol. Thus, REST/NRSF cannot enter the nucleus and bind to the 21 base-pair RE-1/NRSE regulatory element. An adequate repression of specific target genes are of fundamental importance, as many are involved in the proper development of neuronal receptors, neurotransmitters, synaptic vesicle proteins, and channel proteins. A deficiency in the proper development of these proteins can cause the neural dysfunctions seen in Huntington's disease. In addition to the lack of repression due to the inactive REST/NRSF, mutated huntingtin protein can also decrease the transcription of the brain-derived neurotropic factor (BDNF) gene. BDNF influences the survival and development of neurons in the central nervous system as well as the peripheral nervous system. This abnormal repression occurs when the RE1/NRSE region within the BDNF promoter region is activated by the binding of REST/NRSF, which leads to the lack of transcription of the BDNF gene.[12] Hence, the anomalous repression of the BDNF protein suggests a significant impact in Huntington's disease.
Current research on REST/NRSF and ventricular hypertrophy in mammals
REST/NRSF in conjunction with RE1/NRSE also acts outside the nervous system as regulators and repressors. Current research has linked RE1/NRSE activity with the regulation of the expression of the atrial natriuretic peptide (ANP) gene.[10] An NRSE regulatory region is present in the 3’ untranslated region of the ANP gene and acts as a mediator for its appropriate expression. The protein encoded by the ANP gene is important during embryonic development for the maturation and development of cardiac myocytes. However, during early childhood and throughout adulthood, ANP expression is suppressed or kept to a minimum in the ventricle. Thus, an abnormal induction of the ANP gene can lead to ventricular hypertrophy and severe cardiac consequences. In order to maintain the repression of the gene, NRSF (neuron-restrictive silencer factor) or REST binds to the NRSE region in the 3’untranslated region of the ANP gene. Furthermore, the NRSF-NRSE complex recruits a transcriptional corepressor known as mSin3.[10] This leads to the activity of histone deacetylase in the region and the repression of the gene. Therefore, studies have revealed the correlation between REST/NRSF and RE1/NRSE in regulating the ANP gene expression in ventricular myocytes. A mutation in either the NRSF or NRSE can lead to an undesirable development of ventricular myocytes, due to lack of repression, which can then cause ventricular hypertrophy. Left ventricular hypertrophy, for example, increases an individual’s chance of sudden death due to a ventricular arrhythmia resulting from the increased ventricular mass.[13] In addition to the influence on the ANP gene, the NRSE sequence regulates other cardiac embryonic genes, such as brain natriuretic peptide BNP, skeletal α-actin, and Na, K – ATPase α3 subunit.[10] Hence, the regulatory activity of both NRSE and NRSF in mammals prevents not only neural dysfunctions but also physiological and phenotypical abnormalities in other non-neuronal regions of the body.

Mutations in polycomb-group response elements (PREs)
The Polycomb-group (PcG) regulatory complexes are known for their influence in the epigenetic regulation of stem cells, especifically in hematopoietic stem cells. The Polycomb Repressive Complex 1 (PRC 1) is directly involved in the process of hematopoiesis, and functions together with, for example, the PcG gene “Bmi1”. Studies in mice indicate that organisms with mutated “Bmi1” demonstrate deficient mitochondrial functioning, and also hindered the ability of hematopoietic cells to self-renew. Likewise, mutations in PRC2 genes were related to hematological conditions such as acute lymphoblastic leukemia (ALL), which is a form of leukemia. Hence, Polycomb-group genes and proteins are involved in the proper maintenance of hematopoiesis in the body.[14]
References
- 1 2 3 4 5 6 Maston, Glenn; Sarah Evans; Michael Green (23 May 2006). "Transcriptional regulatory elements in the Human Genome" (PDF). Annual Reviews. Retrieved 2 April 2013.
- 1 2 3 4 5 6 7 Ogbourne, Steven; Toni Antalis (1998). "Transcriptional control and the role of silencers in transcriptional regulation in eukaryotes" (PDF). Biochem. J. 331 (1): 1–14. doi:10.1042/bj3310001. PMC 1219314. PMID 9512455. Retrieved 2 April 2013.
- 1 2 "Control of Genetic Systems in Prokaryotes and Eukaryotes". University of Illinois at Chicago. Retrieved 2 April 2013.
- 1 2 "Eukaryotic Gene Control". Kenyon College. Retrieved 1 April 2013.
- ↑ "Gene Regulation in Eukaryotes". Eastern Michigan University. Retrieved 7 April 2013.
- 1 2 3 "Gene Regulation in Eukaryotes". Kimball's Biology Pages. Retrieved 7 April 2013.
- ↑ Brown, TA (2002). Genomes. Oxford: Wiley-Liss.
- ↑ Schoenherr, CJ; Anderson DJ (3 March 1995). "The neuron-restrictive silencer factor (NRSF): a coordinate repressor of multiple neuron-specific genes.". Science 267 (5202): 1360–3. doi:10.1126/science.7871435. PMID 7871435.
- 1 2 3 4 Olguín, Patricio; Pablo Ote ́ıza, Eduardo Gamboa, Jose ́ Luis Go ́mez-Ska ́rmeta, and Manuel Kukuljan (8 March 2006). "RE-1 Silencer of Transcription/Neural Restrictive Silencer Factor Modulates Ectodermal Patterning during Xenopus Development" (PDF). The Journal of Neuroscience. Retrieved 3 April 2013. Cite uses deprecated parameter
|coauthors=(help) - 1 2 3 4 Kuwahara, Koichiro; Yoshihiko Saito, Emiko Ogawa, Nobuki Takahashi, Yasuaki Nakagawa, Yoshihisa Naruse, Masaki Harada, Ichiro Hamanaka, Takehiko Izumi, Yoshihiro Miyamoto, Ichiro Kishimoto, Rika Kawakami, Michio Nakanishi, Nozomu Mori, and Kazuwa Nakao (21 March 2001). "The Neuron-Restrictive Silencer Element–Neuron-Restrictive Silencer Factor System Regulates Basal and Endothelin 1-Inducible Atrial Natriuretic Peptide Gene Expression in Ventricular Myocytes". Molecular and Cellular Biology 21 (6): 2085–97. doi:10.1128/MCB.21.6.2085-2097.2001. PMC 86819. PMID 11238943. Retrieved 21 March 2013. Cite uses deprecated parameter
|coauthors=(help) - ↑ Walker, FO (20 January 2007). "Huntington's disease.". Lancet 369 (9557): 218–28. doi:10.1016/S0140-6736(07)60111-1. PMID 17240289.
- ↑ Zuccato, C; Belyaev N; Conforti P; Ooi L; Tartari M; Papadimou E; MacDonald M; Fossale E; Zeitlin S; Buckley N; Cattaneo E. (27 June 2007). "Widespread disruption of repressor element-1 silencing transcription factor/neuron-restrictive silencer factor occupancy at its target genes in Huntington's disease.". The Journal of Neuroscience. Retrieved 21 March 2013.
- ↑ Rials, Seth; Ying Wu, MD; Nancy Ford, BS; Ferrel J. Pauletto, MD; Sandra V. Abramson, MD; Andrew M. Rubin, MD; Roger A. Marinchak, MD; Peter R. Kowey, MD (1995). "Effect of Left Ventricular Hypertrophy and Its Regression on Ventricular Electrophysiology and Vulnerability to Inducible Arrhythmia in the Feline Heart". American Heart Association. Retrieved 3 April 2013. Cite uses deprecated parameter
|coauthors=(help) - ↑ Sashida, Goro; Iwama, Atsushi. "Epigenetic regulation of hematopoiesis". International Journal of Hematology 96 (4): 405–412. doi:10.1007/s12185-012-1183-x.
External links
- Silencer Elements at the US National Library of Medicine Medical Subject Headings (MeSH)
| ||||||||||||||||||||||||||||||||||||||||||