Small nuclear ribonucleoprotein polypeptide A
| Small nuclear ribonucleoprotein polypeptide A |
|---|
PDB rendering based on 1aud. |
| Available structures |
| PDB |
Ortholog search: PDBe, RCSB |
| List of PDB id codes |
|
1AUD, 1CX0, 1DRZ, 1DZ5, 1FHT, 1M5K, 1M5O, 1M5P, 1M5V, 1NU4, 1OIA, 1SJ3, 1SJ4, 1SJF, 1U6B, 1URN, 1VBX, 1VBY, 1VBZ, 1VC0, 1VC5, 1VC6, 1VC7, 1ZZN, 2A3J, 2NZ4, 2OIH, 2OJ3, 2U1A, 3BO2, 3BO3, 3BO4, 3CUL, 3CUN, 3EGZ, 3G8S, 3G8T, 3G96, 3G9C, 3HHN, 3IIN, 3IRW, 3IWN, 3K0J, 3L3C, 3MUM, 3MUR, 3MUT, 3MUV, 3MXH, 3P49, 3PGW, 3R1H, 3R1L, 3UCU, 3UCZ, 3UD3, 3UD4, 3UTR, 4C4W
|
|
|
| Identifiers |
|---|
| Symbols | SNRPA ; Mud1; U1-A; U1A |
|---|
| External IDs | OMIM: 182285 MGI: 1855690 HomoloGene: 3380 GeneCards: SNRPA Gene |
|---|
|
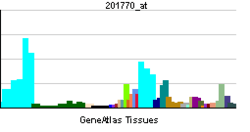
| RNA expression pattern |
|---|
|
| More reference expression data |
| Orthologs |
|---|
| Species | Human | Mouse | |
|---|
| Entrez | 6626 | 53607 | |
|---|
| Ensembl | ENSG00000077312 | ENSMUSG00000061479 | |
|---|
| UniProt | P09012 | Q62189 | |
|---|
| RefSeq (mRNA) | NM_004596 | NM_001046637 | |
|---|
| RefSeq (protein) | NP_004587 | NP_001040102 | |
|---|
| Location (UCSC) | Chr 19:
41.26 – 41.27 Mb | Chr 7:
27.19 – 27.2 Mb | |
|---|
| PubMed search | | | |
|---|
|
U1 small nuclear ribonucleoprotein A is a protein that in humans is encoded by the SNRPA gene.[1][2]
Interactions
Small nuclear ribonucleoprotein polypeptide A has been shown to interact with CDC5L.[3]
References
- ↑ Schonk D, van Dijk P, Riegmann P, Trapman J, Holm C, Willcocks TC, Sillekens P, van Venrooij W, Wimmer E, Geurts van Kessel A et al. (January 1991). "Assignment of seven genes to distinct intervals on the midportion of human chromosome 19q surrounding the myotonic dystrophy gene region". Cytogenet Cell Genet 54 (1-2): 15–9. doi:10.1159/000132946. PMID 1701111.
- ↑ "Entrez Gene: SNRPA small nuclear ribonucleoprotein polypeptide A".
- ↑ Ajuh, P; Kuster B; Panov K; Zomerdijk J C; Mann M; Lamond A I (December 2000). "Functional analysis of the human CDC5L complex and identification of its components by mass spectrometry". EMBO J. (ENGLAND) 19 (23): 6569–81. doi:10.1093/emboj/19.23.6569. ISSN 0261-4189. PMC 305846. PMID 11101529.
Further reading
- Hoffman DW, Query CC, Golden BL et al. (1991). "RNA-binding domain of the A protein component of the U1 small nuclear ribonucleoprotein analyzed by NMR spectroscopy is structurally similar to ribosomal proteins.". Proc. Natl. Acad. Sci. U.S.A. 88 (6): 2495–9. doi:10.1073/pnas.88.6.2495. PMC 51259. PMID 1826055.
- Nelissen RL, Sillekens PT, Beijer RP et al. (1991). "Structure, chromosomal localization and evolutionary conservation of the gene encoding human U1 snRNP-specific A protein.". Gene 102 (2): 189–96. doi:10.1016/0378-1119(91)90077-O. PMID 1831431.
- Jessen TH, Oubridge C, Teo CH et al. (1991). "Identification of molecular contacts between the U1 A small nuclear ribonucleoprotein and U1 RNA.". EMBO J. 10 (11): 3447–56. PMC 453073. PMID 1833186.
- Nagai K, Oubridge C, Jessen TH et al. (1991). "Crystal structure of the RNA-binding domain of the U1 small nuclear ribonucleoprotein A.". Nature 348 (6301): 515–20. doi:10.1038/348515a0. PMID 2147232.
- Sillekens PT, Habets WJ, Beijer RP, van Venrooij WJ (1988). "cDNA cloning of the human U1 snRNA-associated A protein: extensive homology between U1 and U2 snRNP-specific proteins.". EMBO J. 6 (12): 3841–8. PMC 553857. PMID 2962859.
- Oubridge C, Ito N, Evans PR et al. (1994). "Crystal structure at 1.92 A resolution of the RNA-binding domain of the U1A spliceosomal protein complexed with an RNA hairpin.". Nature 372 (6505): 432–8. doi:10.1038/372432a0. PMID 7984237.
- Howe PW, Nagai K, Neuhaus D, Varani G (1994). "NMR studies of U1 snRNA recognition by the N-terminal RNP domain of the human U1A protein.". EMBO J. 13 (16): 3873–81. PMC 395300. PMID 8070414.
- Allain FH, Gubser CC, Howe PW et al. (1996). "Specificity of ribonucleoprotein interaction determined by RNA folding during complex formulation.". Nature 380 (6575): 646–50. doi:10.1038/380646a0. PMID 8602269.
- Avis JM, Allain FH, Howe PW et al. (1996). "Solution structure of the N-terminal RNP domain of U1A protein: the role of C-terminal residues in structure stability and RNA binding.". J. Mol. Biol. 257 (2): 398–411. doi:10.1006/jmbi.1996.0171. PMID 8609632.
- Lu J, Hall KB (1997). "Tertiary structure of RBD2 and backbone dynamics of RBD1 and RBD2 of the human U1A protein determined by NMR spectroscopy.". Biochemistry 36 (34): 10393–405. doi:10.1021/bi9709811. PMID 9265619.
- Allain FH, Howe PW, Neuhaus D, Varani G (1997). "Structural basis of the RNA-binding specificity of human U1A protein.". EMBO J. 16 (18): 5764–72. doi:10.1093/emboj/16.18.5764. PMC 1170207. PMID 9312034.
- Neubauer G, King A, Rappsilber J et al. (1998). "Mass spectrometry and EST-database searching allows characterization of the multi-protein spliceosome complex.". Nat. Genet. 20 (1): 46–50. doi:10.1038/1700. PMID 9731529.
- Ferré-D'Amaré AR, Zhou K, Doudna JA (1998). "Crystal structure of a hepatitis delta virus ribozyme.". Nature 395 (6702): 567–74. doi:10.1038/26912. PMID 9783582.
- Lutz CS, Cooke C, O'Connor JP et al. (1998). "The snRNP-free U1A (SF-A) complex(es): identification of the largest subunit as PSF, the polypyrimidine-tract binding protein-associated splicing factor.". RNA 4 (12): 1493–9. doi:10.1017/S1355838298981183. PMC 1369720. PMID 9848648.
- Hetzer M, Mattaj IW (2000). "An ATP-dependent, Ran-independent mechanism for nuclear import of the U1A and U2B" spliceosome proteins.". J. Cell Biol. 148 (2): 293–303. doi:10.1083/jcb.148.2.293. PMC 2174293. PMID 10648562.
- Klein Gunnewiek JM, Hussein RI, van Aarssen Y et al. (2000). "Fourteen residues of the U1 snRNP-specific U1A protein are required for homodimerization, cooperative RNA binding, and inhibition of polyadenylation.". Mol. Cell. Biol. 20 (6): 2209–17. doi:10.1128/MCB.20.6.2209-2217.2000. PMC 110837. PMID 10688667.
- Varani L, Gunderson SI, Mattaj IW et al. (2000). "The NMR structure of the 38 kDa U1A protein - PIE RNA complex reveals the basis of cooperativity in regulation of polyadenylation by human U1A protein.". Nat. Struct. Biol. 7 (4): 329–35. doi:10.1038/74101. PMID 10742179.
- Ajuh P, Kuster B, Panov K et al. (2001). "Functional analysis of the human CDC5L complex and identification of its components by mass spectrometry.". EMBO J. 19 (23): 6569–81. doi:10.1093/emboj/19.23.6569. PMC 305846. PMID 11101529.
- Simmons A, Aluvihare V, McMichael A (2001). "Nef triggers a transcriptional program in T cells imitating single-signal T cell activation and inducing HIV virulence mediators.". Immunity 14 (6): 763–77. doi:10.1016/S1074-7613(01)00158-3. PMID 11420046.
PDB gallery |
|---|
| | 1aud: U1A-UTRRNA, NMR, 31 STRUCTURES |
| 1cx0: HEPATITIS DELTA VIRUS RIBOZYME |
| 1drz: U1A SPLICEOSOMAL PROTEIN/HEPATITIS DELTA VIRUS GENOMIC RIBOZYME COMPLEX |
| 1dz5: THE NMR STRUCTURE OF THE 38KDA U1A PROTEIN-PIE RNA COMPLEX REVEALS THE BASIS OF COOPERATIVITY IN REGULATION OF POLYADENYLATION BY HUMAN U1A PROTEIN |
| 1fht: RNA-BINDING DOMAIN OF THE U1A SPLICEOSOMAL PROTEIN U1A117, NMR, 43 STRUCTURES |
| 1m5k: Crystal structure of a hairpin ribozyme in the catalytically-active conformation |
| 1m5o: Transition State Stabilization by a Catalytic RNA |
| 1m5p: Transition State Stabilization by a Catalytic RNA |
| 1m5v: Transition State Stabilization by a Catalytic RNA |
| 1nu4: U1A RNA binding domain at 1.8 angstrom resolution reveals a pre-organized C-terminal helix |
| 1oia: U1A RNP DOMAIN 1-95 |
| 1sj3: Hepatitis Delta Virus Gemonic Ribozyme Precursor, with Mg2+ Bound |
| 1sj4: Crystal structure of a C75U mutant Hepatitis Delta Virus ribozyme precursor, in Cu2+ solution |
| 1sjf: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, in Cobalt Hexammine solution |
| 1u6b: CRYSTAL STRUCTURE OF A SELF-SPLICING GROUP I INTRON WITH BOTH EXONS |
| 1urn: U1A MUTANT/RNA COMPLEX + GLYCEROL |
| 1vbx: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, in EDTA solution |
| 1vby: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, and Mn2+ bound |
| 1vbz: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, in Ba2+ solution |
| 1vc0: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, in Imidazole and Sr2+ solution |
| 1vc5: Crystal Structure of the Wild Type Hepatitis Delta Virus Gemonic Ribozyme Precursor, in EDTA solution |
| 1vc6: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Product with C75U Mutaion, cleaved in Imidazole and Mg2+ solutions |
| 1vc7: Crystal Structure of the Hepatitis Delta Virus Gemonic Ribozyme Precursor, with C75U mutaion, in Sr2+ solution |
| 1zzn: Crystal structure of a group I intron/two exon complex that includes all catalytic metal ion ligands. |
| 2nz4: Structural investigation of the GlmS ribozyme bound to its catalytic cofactor |
| 2oih: Hepatitis Delta Virus gemonic ribozyme precursor with C75U mutation and bound to monovalent cation Tl+ |
| 2oj3: Hepatitis Delta Virus ribozyme precursor structure, with C75U mutation, bound to Tl+ and cobalt hexammine (Co(NH3)63+) |
| 2u1a: RNA BINDING DOMAIN 2 OF HUMAN U1A PROTEIN, NMR, 20 STRUCTURES |
|
|
|
|
|---|
| | snRNP | |
|---|
| | hnRNP | |
|---|
| | Other transcription | |
|---|
| | Translation | |
|---|
| Vault ribonucleoprotein
particles | |
|---|
| | Other | |
|---|
| Index of genetics |
|---|
| | Description |
- Gene expression
- DNA
- replication
- cycle
- recombination
- repair
- binding proteins
- Transcription
- factors
- regulators
- nucleic acids
- RNA
- RNA binding proteins
- ribonucleoproteins
- repeated sequence
- modification
- Translation
- ribosome
- modification
- nexins
- Proteins
- domains
- Structure
- primary
- secondary
- tertiary
- quaternary
|
|---|
| | Disease |
- Replication and repair
- Transcription factor
- Transcription
- Translation
|
|---|
|
|