Mad1
| Mad1 | |
|---|---|
 | |
| Crystal structure, tetramer of Mad1-Mad2 complex, yellow and red=Mad1 monomers, palegreen= Mad2 monomers | |
| Identifiers | |
| Organism | |
| Symbol | MAD1 |
| Entrez | 852794 |
| PDB | 1GO4 |
| RefSeq (mRNA) | NM_001180951.3 |
| RefSeq (Prot) | NP_011429.3 |
| UniProt | P40957 |
| Other data | |
| Chromosome | VII: 0.35 - 0.35 Mb |
Mad1 is a non-essential protein in yeast which has a function in the spindle assembly checkpoint (SAC).[1] This checkpoint monitors chromosome attachment to spindle microtubules and prevents cells from starting anaphase until the spindle is built up. The name Mad refers to the observation that mutant cells are mitotic arrest deficient (MAD) during microtubule depolymerization. Mad1 recruits the anaphase inhibitor Mad2 to unattached kinetochores and is essential for Mad2-Cdc20 complex formation in vivo but not in vitro. In vivo, Mad1 acts as a competitive inhibitor of the Mad2-Cdc20 complex.[2] Mad1 is phosphorylated by Mps1 which then leads together with other activities to the formation of the mitotic checkpoint complex (MCC). Thereby it inhibits the activity of the anaphase-promoting complex/cyclosome (APC/C). Homolog’s of Mad1 are conserved in eukaryotes from yeast to mammals.
Introduction
In the early 90s, yeast genes were identified which mutations resulted in a defect in mitotic arrest in response to microtubule disassembly (mitotic arrest deficient genes - MAD genes). This cells showed during division no mitotic arrest in the presence of microtubule polymerization inhibitors and were therefore not able to delay cell division.[1] The genes identified included the MAD1, MAD2 and MAD3 genes. They are conserved in all eukaryotes and are involved in a pathway that is active in prometaphase to prevent the premature separation of sister chromatids and constitute the so-called spindle assembly checkpoint(SAC). This checkpoint monitors the status of chromosome attachment to the mitotic spindle and inhibits the metaphase to anaphase transition by preventing the activation of the anaphase-promoting complex/cyclosome (APC/C), and thereby the degradation of cell cycle regulators.[3] Mad1 is in this pathway accumulated at unattached kinetochores and acts as a sensor for unattached kinetochores in this machinery.
Function
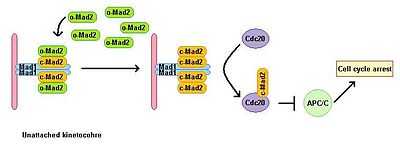
Eukaryotic cells show a mitotic arrest in the presence of microtubule polymerization inhibitors. A spindle assembly checkpoint monitors the status of the spindle and links the metaphase-anaphase transition to proper bipolar attachment of all kinetochores to the mitotic spindle. The spindle assembly checkpoint inhibits the activity of the anaphase promoting complex by preventing degradation of downstream effectors, which otherwise lead to anaphase onset and exit from mitosis. Depletion of Mad1 leads to the loss of SAC function. Mad1 localises predominantly at unattached kinetochores and triggers mitotic arrest in case of a single unattached kinetochore. Mad1 recruits the important SAC component Mad2 to unattached kinetochores (Fig. 2) and induces mitotic arrest signal amplification. There is a pool of free cytoplasmic Mad2 in its inactive open conformation called o-MAD2. When bound to Mad1, Mad2 adopts an active conformation called closed (c-Mad2) and forms a heterotetramer of two Mad1 and two c-Mad2 units. The heterotetramer of Mad1–c-Mad2 is very stable and works as a catalytic receptor for free cytoplasmic o-Mad2. Free o-Mad2 binds to this receptor and changes its conformation to the active closed form. This second c-MAD2 is transferred to Cdc20 with yet unknown mechanism and forms Cdc20–c-Mad2 complex. This complex is an essential component of mitotic checkpoint complex (MCC). MCC binds and inhibits APC/C and therefore arrests progression through mitosis.[3][4]
Regulation
There are two upstream checkpoint kinases implicated in regulating Mad1 function through phosphorylation.[5] Mps1 phosphorylates Mad1 both in vitro and in vivo and is thought to regulate Mad1 and Mad2 localization to kinetochores and their interaction dynamics. BUB1 is the other kinase that recruits Mad1 to kinetochores and activates it if a kinetochore is unattached.[3] If a kinetochore is attached to spindle, SAC inhibitor p31comet inhibits Mad1 mediated conformational rearrangement of Mad2 and prevents Mad2 from binding to Cdc20.[3]
Structural Features and Mechanism
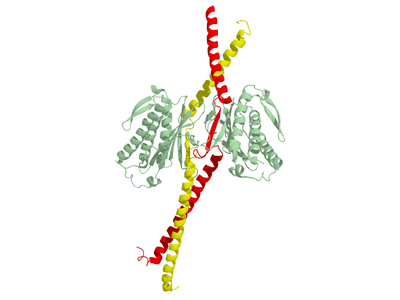
By biochemical methods Mad1 was predicted to encode a 90kD, 718-residue,[6] coiled-coil protein with a characteristic rod shape[1] in 1995. Crystal structures followed soon. Then in 2002 the crystal structure of human Mad1 in complex with human Mad2 forming a tetramer was published (Fig. 1). Due to experimental limitations the structure only shows Mad1 residues 484 - 584. Elongated Mad1 monomers are tightly held together by a parallel coiled-coil involving the N-terminal alpha helices. The Mad1 chains point away from the coiled-coil towards their Mad2 ligands forming two sub-complexes with Mad2. The segment between alpha helices 1 and 2 contains the Mad2 binding domain (Fig. 3). The first part of this binding domain is flexible and adopts different conformations giving rise to an asymmetric complex. In their work, employing thermodynamic studies, Sironi et al.[2] show that Mad1 functions such as to slow down the rate of Mad2-Cdc20 complex formation and therefore acts as a competitive inhibitor in vivo. Furthermore the authors suggest, the Mad1-Mad2 binding sites are buried inside the structure perhaps rendering the binding sites inaccessible for Cdc20 binding. Mad1-Mad2 binding is unusual in that the Mad2 C-terminal folds over Mad1. The authors therefore conclude that an unperturbed Mad1-Mad2 complex will not release Mad2 requiring a novel, so far poorly understood, mechanism of conformational change.[2]
Cancer
Mismatches in chromosome number (aneuploidies) during meiosis are responsible for human diseases like Down's syndrome and also emerge frequently in cancer cells. The essential function of SAC gives rise to the hypothesis that mutations of the SAC and especially inactivation of SAC might be a reason for tumorigenesis or at least facilitate tumorigenesis.[3] Against this idea, it was shown that cancer cells undergo apoptosis when components of the SAC are not present.[7] In this model, in contrast to the other model, SAC inactivation becomes a potential way to kill rapidly dividing cancer cells. The molecular links between Mad1p, the SAC, apoptosis and cancer are still not fully understood.[3]
See also
- MAD2
- Hyperphosphorylation
References
- ↑ 1.0 1.1 1.2 Hardwick K, Murray A (1995). "Mad1p, a phosphoprotein component of the spindle assembly checkpoint in budding yeast". The Journal of Cell Biology 131 (3): 709–720. doi:10.1083/jcb.131.3.709. PMC 2120625. PMID 7593191.
- ↑ 2.0 2.1 2.2 Sironi L, Mapelli M, Knapp S, De Antoni A, Jeang K, Musacchio A (2002). "Crystal structure of the tetrameric Mad1–Mad2 core complex: implications of a 'safety belt' binding mechanism for the spindle checkpoint". EMBO 21 (10): 2496–2506. doi:10.1093/emboj/21.10.2496. PMC 126000. PMID 12006501.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 Musacchio A, Salmon E (May 2007). "The spindle-assembly checkpoint in space and time". Nat. Rev. Mol. Cell Biol. 8 (5): 379–93. doi:10.1038/nrm2163. PMID 17426725.
- ↑ Yu H (Apr 2006). "Structural activation of Mad2 in the mitotic spindle checkpoint: the two-state Mad2 model versus the Mad2 template model". J Cell Biol. 173 (2): 153–157. doi:10.1083/jcb.200601172. PMC 2063805. PMID 16636141.
- ↑ Bharadwaj R, Yu H (2000). "The spindle checkpoint, aneuploidy, and cancer". Oncogene 23 (11): 2016–27. doi:10.1038/sj.onc.1207374. PMID 15021889.
- ↑ Chen R, Shevchenko A, Mann M, Murray A (1998). "Spindle Checkpoint Protein Xmad1 Recruits Xmad2 to Unattached Kinetochores". The Journal of Cell Biology 134 (2): 283–295. doi:10.1083/jcb.143.2.283. PMC 2132829. PMID 9786942.
- ↑ Kops G, Foltz D, Cleveland D (June 2004). "Lethality to human cancer cells through massive chromosome loss by inhibition of the mitotic checkpoint". Proc. Natl. Acad. Sci. U.S.A. 101 (23): 8699–704. doi:10.1073/pnas.0401142101. PMC 423258. PMID 15159543.