Reverse transcriptase
| Reverse Transcriptase | |
|---|---|
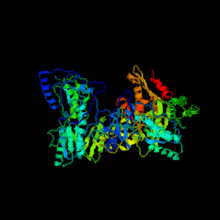 3D model of HIV reverse transcriptase |
|
| Other names: | Deoxynucleoside-triphosphate: DNA deoxynucleotidyltransferase (RNA-directed)
|
| Database Links | |
| EC number: | 2.7.7.49 |
In biochemistry, a reverse transcriptase, also known as RNA-dependent DNA polymerase, is a DNA polymerase enzyme that transcribes single-stranded RNA into double-stranded DNA. It also helps in the formation of a double helix DNA once the RNA has been reverse transcribed into a single strand cDNA. Normal transcription involves the synthesis of RNA from DNA; hence, reverse transcription is the reverse of this.
Reverse transcriptase was discovered by Howard Temin at the University of Wisconsin-Madison, and independently by David Baltimore in 1970 at MIT. The two shared the 1975 Nobel Prize in Physiology or Medicine with Renato Dulbecco for their discovery.
Well studied reverse transcriptases include:
- HIV-1 reverse transcriptase from human immunodeficiency virus type 1 (PDB 1HMV)
- M-MLV reverse transcriptase from the Moloney murine leukemia virus
- AMV reverse transcriptase from the avian myeloblastosis virus
- Telomerase reverse transcriptase that maintains the telomeres of eukaryotic chromosomes
Contents |
Function
Viruses
The enzyme is encoded and used by reverse-transcribing viruses, which use the enzyme during the process of replication. Reverse-transcribing RNA viruses, such as retroviruses, use the enzyme to reverse-transcribe their RNA genomes into DNA, which is then integrated into the host genome and replicated along with it. Reverse-transcribing DNA viruses, such as the hepadnaviruses, can allow RNA to serve as a template in assembling, and making DNA strands. HIV infects humans with the use of this enzyme. Without reverse transcriptase, the viral genome would not be able to incorporate into the host cell, resulting in the failure of the ability to replicate. Unlike bacteria, retroviruses use preexisting host-encoded transfer RNAs as primers.
Process of Reverse Transcription
Retroviral RNA is arranged in 5’ terminus to 3’ terminus. The site where the primer is annealed to viral RNA is called the primer-binding site (PBS). The RNA 5’end to the PBS site is called U5, and the RNA 3’ end to the PBS is called the leader. The tRNA primer is unwound between 14 and 22 nucleotides and forms a base-paired duplex with the viral RNA at PBS. PBS locates near the 5’ terminus of viral RNA is unusual because reverse transcriptase synthesize DNA from 3’ end of the primer in the 5’ to 3’ direction. Therefore, the primer and reverse transcriptase must be relocated to 3’ end of viral RNA. In order to accomplish this reposition, multiple steps and various enzymes included DNA polymerase, ribonuclease H(RNase H) and polynucleotide unwinding are needed. [1]
The reverse transcription of retroviral RNA is a discontinuous process. The production of DNA begins with short and discrete segments which will be elongated during a process called strand transfer. Strand transfer involves translocation of short DNA product from initial synthesis site to acceptor template regions at the other end of the genome. Thus, the final product of reverse transcriptase is a linear double-stranded DNA.[2]
Eukaryotes
Self-replicating stretches of eukaryotic genomes known as retrotransposons utilize reverse transcriptase to move from one position in the genome to another via a RNA intermediate. They are found abundantly in the genomes of plants and animals. Telomerase is another reverse transcriptase found in many eukaryotes, including humans, which carries its own RNA template; this RNA is used as a template for DNA replication[3].
Prokaryotes
Reverse transcriptases are also found in bacterial Retron msr RNAs, distinct sequences which code for reverse transcriptase, and are used in the synthesis of msDNA. In order to initiate synthesis of DNA, a primer is needed. In bacteria, the primer is synthesized during replication.[4]
Structure
Reverse transcriptase enzymes include an RNA-dependent DNA polymerase and a DNA-dependent DNA polymerase, which work together to perform transcription. In addition to the transcription function, retroviral reverse transcriptases have a domain belonging to the RNase H family which is vital to their replication.
Replication fidelity
There are three different replication systems during the life cycle of a retrovirus. First of all, the reverse transcriptase synthesize viral DNA from viral RNA, and then from newly made complementary DNA strand. The second replication process occurs when host cellular DNA polymerase replicates the integrated viral DNA. Lastly, RNA polymerase II transcribes the proviral DNA into RNA which will be packed into virions. Therefore, mutation can occur during one or all of these replication steps.[5]
Reverse transcriptase has a high error rate when transcribing RNA into DNA since, unlike DNA polymerases, it has no proofreading ability. This high error rate allows mutations to accumulate at an accelerated rate relative to proofread forms of replication. The commercially available reverse transcriptases produced by Promega are quoted by their manuals as having error rates in the range of 1 in 17,000 bases for AMV and 1 in 30,000 bases for M-MLV[6]
Applications
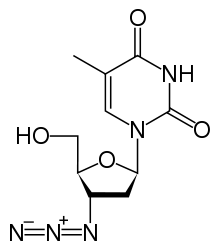
Antiviral drugs
-
For more details on this topic, see Reverse transcriptase inhibitor.
As HIV uses reverse transcriptase to copy its genetic material and generate new viruses (part of a retrovirus proliferation circle), specific drugs have been designed to disrupt the process and thereby suppress its growth. Collectively, these drugs are known as reverse transcriptase inhibitors and include the nucleoside and nucleotide analogues zidovudine (trade name Retrovir), lamivudine (Epivir) and tenofovir (Viread), as well as non-nucleoside inhibitors, such as nevirapine (Viramune).
Molecular biology
-
For more details on this topic, see Reverse transcription polymerase chain reaction.
Reverse transcriptase is commonly used in research to apply the polymerase chain reaction technique to RNA in a technique called reverse transcription polymerase chain reaction (RT-PCR). The classical PCR technique can be applied only to DNA strands, but, with the help of reverse transcriptase, RNA can be transcribed into DNA, thus making PCR analysis of RNA molecules possible. Reverse transcriptase is used also to create cDNA libraries from mRNA. The commercial availability of reverse transcriptase greatly improved knowledge in the area of molecular biology, as, along with other enzymes, it allowed scientists to clone, sequence, and characterise DNA.
See also
- cDNA library
- DNA polymerase
- msDNA
- Reverse transcribing virus
- RNA polymerase
- Telomerase
- Retrotransposon marker
External links
- animation of reverse transcriptase action and three reverse transcriptase inhibitors
- Molecule of the month (September 2002) at the Protein Data Bank
- BRENDA database entry - highly detailed information from a protein database
- MeSH RNA+Transcriptase
- EC 2.7.7.49
- [[1]] - HIV Replication 3D Medical Animation. (Nov 2008). Video by Boehringer Ingelheim.
References
- ↑ Weiss, R., N. Teich, H. Varmus, and J. Coffin (1985). "RNA tumor viruses". in Jonathan, L., Ashock, A., David, C.. Molecular biology of tumor viruses (2nd ed.). New York: Cold Spring Harbor.
- ↑ Telesnitsky, A., Goff, S.P. (1993). "Strong-stop strand transfer during reverse transcription". in Skalka, M. A., Goff, S.P. Reverse transcriptase (1st ed.). New York: Cold Spring Harbor. pp. 49. ISBN 0-87969-382-7.
- ↑ Lodish, et al, Molecular Cell Biology (2004), 5th edn, W. H. Freeman and Company, New York, ISBN 0-7167-4366-3
- ↑ Hurwitz, J. and J. Leis (1972). "Directing influence of DNA in the reaction". RNA-dependent DNA polymerase activity of RNA tumor viruses.
- ↑ Bbenek, K., Kunkel, A. T (1993). "The fidelity of retroviral reverse transcriptases". in Skalka, M. A., Goff, P. S.. Reverse transcriptase. New York: Cold Spring Harbor Laboratory Press. pp. p. 85. ISBN 0-87969-382-7.
- ↑ Promega kit instruction manual (1999)
|
||||||||||||||||||||||||||